Cutting and Joining DNA Molecules - Enabling Techniques
| Home | | Pharmaceutical Microbiology | | Pharmaceutical Microbiology |Chapter: Pharmaceutical Microbiology : Recombinant DNA Technology
DNA isolated from any type of cell can be fragmented using restriction endonucleases. These are enzymes produced by microorganisms which cut foreign DNA and can restrict the proliferation of infecting viruses.
CUTTING AND JOINING DNA MOLECULES
DNA isolated from any type of cell can
be fragmented using restriction endonucleases. These are enzymes produced by
microorganisms which cut foreign DNA and can restrict the proliferation of
infecting viruses. Some of these enzymes cut at specific points known as restriction sites which are palindromic sequences
(complementary sequences with identical nucleotide sequences when read in the
5′ to 3′ direction) of various lengths. For example, EcoRI (Escherichia colirestriction enzyme I) has specificity for
the sequence GAATTC and hydrolyses the G-A phosphodiester bond. Enzymes which
recognize and cut at restriction sites of 4–8 base pairs (bp) are particularly
useful, as the probability of a site appearing in a random DNA fragment is
inversely proportional to its length.
There are two different types of DNA
ends that can be generated using restriction enzymes: cohesive or sticky and
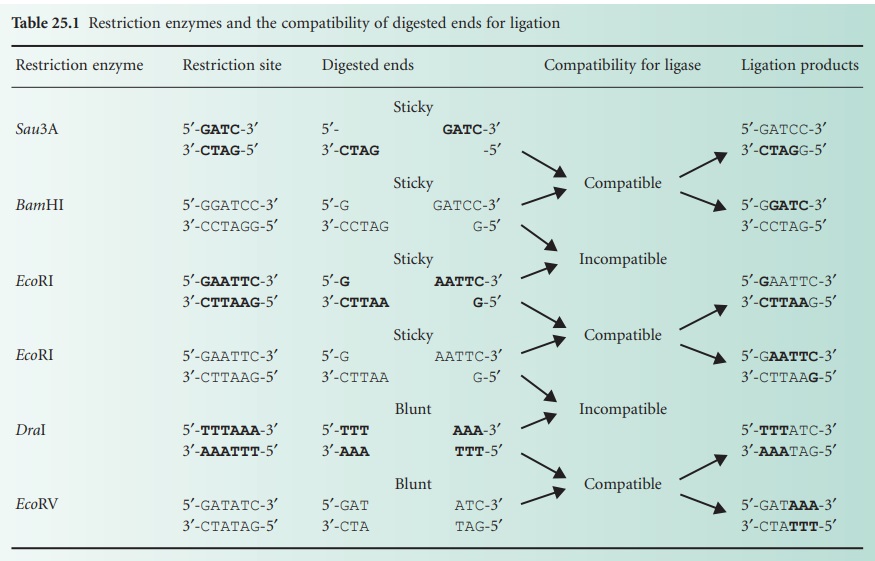
blunt ends (Table 25.1).
DNA fragments obtained by restriction enzyme digestion can be covalently joined
together using the enzyme DNA ligase. There is a limitation, however, with
regards to the type of ends this enzyme is able to bond together. Only blunt
ends generated by some restriction enzymes (e.g. DraI and EcoRV) or compatible
sticky ends generated by either the same restriction enzymes (e.g. Eco RI) or by enzymes that generate complementary
overhanging ends (e.g. Sau3AI and BamHI) will be bonded by the DNA ligase (Table 25.1).
Related Topics
