Bacterial Translation
| Home | | Pharmaceutical Microbiology | | Pharmaceutical Microbiology |Chapter: Pharmaceutical Microbiology : Microbial Genetics and Variations
Bacterial translation may be defined as - ‘the specific process via which the critical nitrogenous-base sequence of mRNA affords determination of the amino acid sequence of protein’.
Bacterial
Translation
Bacterial translation may be
defined as - ‘the specific process via which the critical nitrogenous-base
sequence of mRNA affords determination of the amino acid sequence of protein’.
One may
precisely observe that in an organism that is particularly devoid of a membrane-enclosed nucleus, both ‘transcription’ and ‘translation’ invariably occur in the cytoplasm. Thus, in an eukaryotic organisms, the process of ‘translation’ actually comes into being in a situation when mRNA gains its entry into the cytoplasm.
Figure :
6.7 illustrates the eight major
sequential stages that are intimately involved in the proc-ess of translation, namely :
Stage-1 : Various components that are
essentially required to commence the
‘phenomenon of translation’ first come together.
Stage-2 : On the assembled ribosome a
transfer RNA (tRNA) carrying the ‘first
amino acid’ is duly paired with
the start codon on the mRNA ; and a ‘second amino acid’ being carried by tRNA approaches steadily.
State-3 : Critical place on the chromsome
at which the very first tRNA sites
is known as the P site. Thus, in the
corresponding A site next to it, the
second codon of the mRNA pairs with a tRNA carrying the second
amino acid.
Stage-4 : First amino acid gets
hooked on to the second amino acid
by a peptide linkage 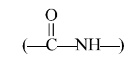 , and the
first tRNA gets released.
, and the
first tRNA gets released.
Note : Nucleotide bases are duly labeled only for
the first two codons.
Stage-5 : Ribosome gradually
moves along the mRNA until the second tRNA is in the P site, and thus the process
continues.
Stage-6 : Ribosome very much
continues to move along the mRNA, and
thus, newer amino acids are
progressively added on to the ‘polypeptide
chain’ strategically.
Stage-7 : Ribosome when
ultimately gets upto the ‘stop codon’, the
duly formed polypeptide is released.
Stage-8 : Last tRNA gets released finally, and thus the ribosome falls apart. Finally, the re-leased polypeptide gives rise to an altogether new protein.
Process of Bacterial Translation : The
various steps encountered in the elaborated process of ‘bacterial translation’ are :
(1) Proteins
are usually synthesized strategically in the 5′ → 3′ direction, as present in DNA and RNA (i.e., nucleic acids).
(2) First
and foremost, the 5′ end of the specific mRNA molecule
becomes associated with a ribosome, which
being the major cellular machinery that
predominantly helps to catalyze the ‘protein synthesis’.
(3) Ribosomal RNA [rRNA] : Ribosomes usually
comprise of two subunits ; of which,
one being a special type of RNA
termed as ribosomal RNA (rRNA) and
the other proteins. At the very
outset of the process of bacterial
translation, the two ribosomal subunits happen to get closer
vis-a-vis the mRNA plus many other components engaged in this phenomenon.
(4) Even
before the suitable amino acids may be joined together to yield a ‘protein’, they should be adequately ‘activated’ by strategic attachment to transfer RNA (tRNA).
Figure : 6.8(a)
represents
the various diagramatic sketch of structures and articulated function of transfer RNA (tRNA).
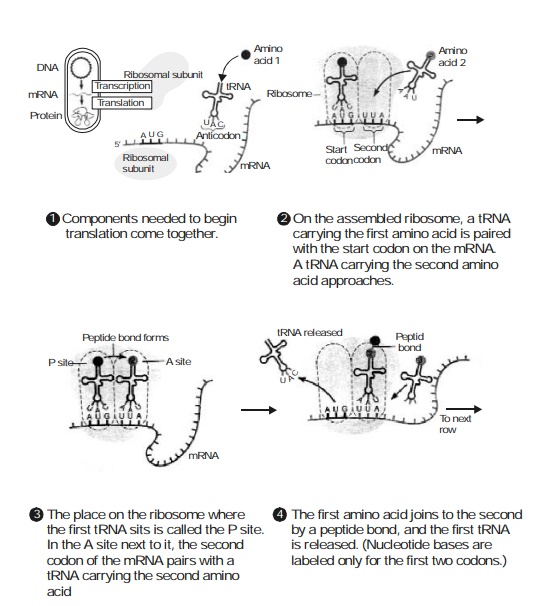
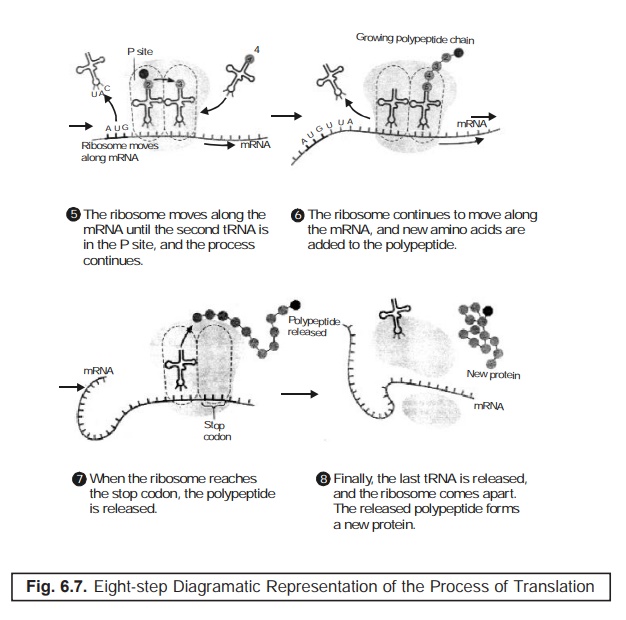
Figure : 6.8(b)
depicts
the manner whereby each different amino acid having a particular tRNA gets duly attached to its
specific tRNA in the course of ‘amino acid activation’ process.
However, this attachment may be adequately achieved by the aid of an amino acid activating enzyme together with sufficient energy derived from adenosine triphosphate (ATP).
Figure : 6.8(c)
illustrates
clearly the way mRNA actually
establishes the precise order wherein amino
acids are duly linked together to give rise to the formation of a protein. Thus, each and every set of three nucleotides of mRNA, usually
termed as codon, evidently specifies
(i.e., codes for) a ‘single amino
acid’.
Example : The following sequence :
AUGCCAGGCAAA
essentially
contains four codons (i.e., four sets of 3 nucleotides of
mRNA) codefying for the amino acids viz.,
methionine (AUG), proline (CCA), glycine (GGC), and lysine
(AAA).
In case,
the bases are grouped in an altogether different manner, the ‘same sequence’ might specify other
amino acids.
Example : AUGC CAG GCA AA
The above
sequence duly encodes : cysteine (UGC),
glutamine (CAG), and alanine (GCA).
Likewise,
AU GCC AGG CAAA
Would
rightly encode alanine (GCC), arginine
(AGG), and glutamine (CAA).
Reading Frames : In fact, all the above cited ‘groupings’ are known as reading frames. Importantly, a
particular reading frame is
invariably determined by the inherent strategic position (status) of the ‘very first codon’ of the gene.
(5) The transfer RNA [tRNA] molecules actually
help to ‘read’ the so called coded message located strategically on
the mRNA.
Anticodon : Anticodon refers to
‘a set of three nucleotides, which is critically positioned on one particular segment of each tRNA molecule, that happens to be complementary to the codon specifically for the ‘amino acid’ being carried by the tRNA [see Fig. 6.8(c)].
(6) It
has been duly observed that in the course of ‘translation’, the highly specific ‘anticodon’ of a molecule of
tRNA gets intimately H-bonded to the complementary
codon strategi-cally located on mRNA.
Example : One may critically observe that a tRNA having the desired anticodon CGA pairs specifically with
the mRNA codon GCU. Therefore, the
eventual pairing of anticodon and codon may usually take place solely at two sites as indicated by the ribosome, such as :
(a) The ‘A’ or ‘aminoacyl-site’,
and
(b) The ‘P’ or ‘peptidyl-site’.
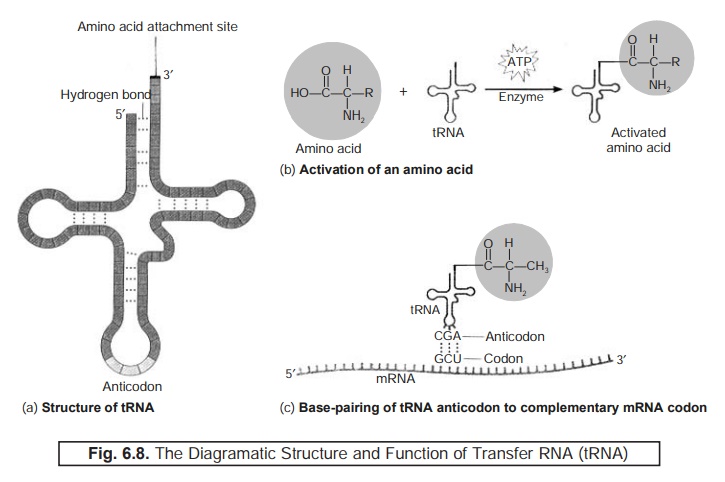
(a) The structure of tRNA is designated in 2D-form.
Each ‘box’ represents a ‘nucleotide’. The critical zones of H-bonding between
‘base pairs’ and ‘loops of unpaired bases’ i.e.,
a typical arrangement to be seen exclusively in RNA molecules.
(b) Activation of ‘each amino acid’ by due
attachment to tRNA.
(c) ‘Anticodon’ by tRNA invariably pairs with its
complementary codon strategically lo-cated on an mRNA strand. The tRNA
displayed specifically carries the amino acid ‘alanine’. The ‘anticodons’ are
mostly represented and duly read in the 5′ → 3′ di-rection ; and, therefore, the anticodon for the
amino acid ‘alanine’ may be read as C–G–A.
[Adapted
From : Tortora GJ et al. : Microbiology : An Introduction,
The
Benjamin and Cummings Publishing Co. Inc., New York, 5th edn., 1995]
Related Topics
