Gene silencing technologies
| Home | | Pharmaceutical Drugs and Dosage | | Pharmaceutical Industrial Management |Chapter: Pharmaceutical Drugs and Dosage: Biotechnology-based drugs
Gene silencing technologies can be exemplified by antisense ODNs, peptide nucleic acids (PNAs), antisense RNA, aptamers, ribozymes, and double-stranded siRNA.
Gene silencing
technologies
Gene
silencing technologies can be exemplified by antisense ODNs, peptide nucleic
acids (PNAs), antisense RNA, aptamers, ribozymes, and double-stranded siRNA.
Antisense oligonucleotides
To
create antisense drugs, nucleotides are linked together in short chains, called
ODNs. When deoxyribonucleotides are
linked in small chains, these are called oligodeoxyribonucleotides.
The sequence of nucleotides in the antisense drugs is complementary to small
segments of mRNA. Each antisense drug is designed to bind to a specific
sequence of nucleotides in its mRNA target to inhibit production of the protein
encoded by the tar-get mRNA. By acting at this earlier stage in the
disease-causing process to prevent the production of a disease-causing protein,
antisense drugs have the potential to provide greater therapeutic benefit than
traditional drugs—which do not act until the disease-causing protein has
already been produced. Antisense drugs also have the potential to be much more
selec-tive or specific than traditional drugs, and therefore more effective,
because they bind to specific mRNA targets through multiple points of
interaction at a single binding site. An oligomer of about 15–20 nucleotides in
length is considered to be the best because this corresponds to both the
appropriate length of a single unique target site in mRNA and the length
required for effective hybridization. Cellular uptake of ODNs occurs by means
of fluid-phase pinocytosis and/or receptor-mediated endocytosis.
Figure 26.2 shows the structures of antisense compounds.
Unmodified ODNs are polyanions with a phosphodiester backbone. They are
rapidly degraded under physiological conditions by enzymes called nucleases,
pri-marily 3'-exonucleases. Because of this, ODN modifications have been
designed to prevent or reduce the rate of degradation. The phosphorothio-ate
modification of the ODN backbone, in which a sulfur atom replaces one of the
nonbridging oxygen atoms in the phosphate group, produces ODNs that are
relatively resistant to cellular and serum nucleases. Methylphosphonate ODNs
have no net charge, which prevents nuclease digestion, but also decreases water
solubility (Figure 26.2a).
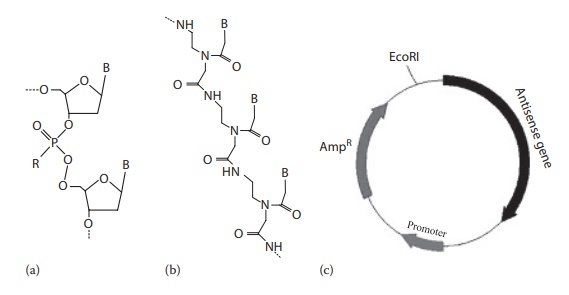
Figure 26.2 Structures of antisense compounds. (a) antisense oligonucleotide (ODN), (b) peptide nucleic acid (PNA), and (c) antisense RNA. R: –O, phospho-diester oligonucleotide; –S, phosphorothioate oligonucleotide; –CH3, methylphosphonate oligonucleotide.
Triplex-forming oligonucleotides
In
contrast to antisense ODNs, triplex-forming oligonucleotides (TFOs) inhibit
gene transcription by forming DNA triple helices in a sequence-specific manner
on polypurine–polypyrimidine tracts. Targeting TFOs to
There are only two copies of targeted gene, whereas there are
thousands of copies of mRNA. Blocking mRNA trans-lation does not prevent the
corresponding gene from being transcribed, which continuously repopulates the
RNA pool. In contrast, prevention of gene transcription can bring down the mRNA
concentration in a more efficient and long-lasting way.
DNA
normally exists in a duplex form (two strands coiled around each other).
However, under some circumstances, DNA can assume triple helix structures.
Triplex helix formation may then prevent the interaction of vari-ous transcription
factors, or it may physically block the initiation or elonga-tion of the
transcription complex. This process is used in the application of TFOs. The
TFOs are specific DNA duplex binding ODNs that can bind to DNA duplex, leading
to triple helix formation and prevention of the transcription process.
Peptide nucleic acids
PNA
has a chemical structure similar to DNA and RNA but differs in the composition
of its backbone. DNA and RNA have a deoxyribose and ribose sugar backbone,
respectively, whereas PNA’s backbone is composed of repeating N-(2-aminoethyl)-glycine units linked by
peptide bonds. The various purine and pyrimidine bases are linked to the
backbone by methylene carbonyl bonds. PNAs are depicted similar as peptides,
with the N-terminus at the first (left) position and the C-terminus at the
right (Figure 26.2b). As the backbone of PNA
contains no charged phosphate groups, the binding between PNA/DNA strands is
stronger than between DNA/DNA strands due to the lack of electrostatic
repulsion. PNAs are resistant to both nucleases and proteases (enzymes that
digest proteins). PNAs can bind to DNA and RNA targets in a sequence-specific
manner to form PNA/DNA and PNA/RNA Watson–Crick double helical structures.
Therefore, PNAs can be used as antisense medicines similar to ODNs.
Antisense RNA
The
antisense mRNA strategy relies on the transfection and subsequent expression of
a plasmid vector whose gene expression cassette carries the cDNA of the gene of
interest subcloned into the vector in an antisense orien-tation (Figure 26.2c). After transfection (the process of introducing foreign genetic material
into cells) into the cells, the plasmid expresses the anti-sense mRNA within
the cell cytoplasm. This antisense mRNA can hybrid-ize exclusively with the mRNA
of the gene of interest and can block protein synthesis (translation). Hence,
antisense mRNA gene medicines require expression vectors and delivery systems
similar to gene therapy medicines (discussed later in this chapter).
MicroRNA
MicroRNAs
(miRNAs) are endogenous, noncoding, small (~22 nucleo-tides) double-stranded
RNAs that participate in gene silencing and post-transcriptional regulation of
gene expression. The miRNAs regulate several cellular processes to impact
overall outcomes in areas such as cell survival and fat metabolism. Each miRNA
has multiple targets and makes global changes in the cellular systems. Changes
in the expression level of miRNAs are associated with phenotypic or performance
differentiation among different cells. For example, high and low titer
producing chinese hamster ovary (CHO) cell lines have shown differences in the
concentra-tions of specific miRNAs. Manipulating the levels of miRNAs in CHO
cells leads to increased recombinant protein production by influencing cell proliferation,
increasing cellular resistance to apoptosis, and increasing specific
productivity.
Gene
medicines that utilize the miRNA pathway might either be exog-enously
administered miRNAs or gene silencing therapies that block the production of
endogenous miRNAs.
Aptamers
Aptamers
are single-stranded or double-stranded nucleic acids that can bind proteins
involved in the regulation and expression of genes (i.e., tran-scription
factors). In addition, they also bind to proteins that perform other regulatory
functions. For example, a 15-mer (i.e., 15 nucleotide long) DNA aptamer binds
to human thrombin and prevents thrombin-catalyzed blood coagulation. In this
approach, the target site is extracellular, and hence the aptamer nucleic acid
does not have to cross the cell membrane to be effec-tive, after parenteral
administration.
Ribozymes
Ribozyme,
also known as RNA enzyme or catalytic RNA, is an RNA mol-ecule having catalytic
enzyme activity that uses either transesterification or a hydrolysis mechanism
to cleave a unique phosphodiester bond in a single-stranded RNA molecule in a
sequence-dependent manner. This process, therefore, leads to interference with
the translation process.
RNA interference
RNAi
is a phenomenon in which double-stranded RNA (dsRNA) mole-cules efficiently and
specifically inhibit gene expression at a posttranscrip-tional level.
Endogenous mRNA exists as a single strand. Mammalian cells have a specific
enzyme, called Dicer, which recognizes the double-stranded RNA and chops it up
into small fragments of between 21–25 base pairs in length. Such a short
double-stranded RNA fragment is called siRNA. The siRNA can bind certain
cellular proteins to form the RNA-induced silenc-ing complex (RISC). The RISC
gets activated when the siRNA unwinds. The activated complex binds to the mRNA
corresponding to the antisense RNA. Thus, siRNA silences a target gene by
binding to its complementary mRNA and by triggering its degradation. The
mechanisms of RNAi are illustrated in Figure 26.3.
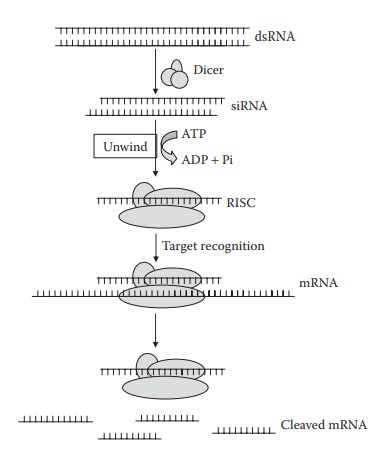
Figure 26.3 Mechanisms of RNA interference. Long double-stranded RNA is cleaved by Dicer into fragments of 21–23 nucleotide siRNAs. Following unwinding, the antisense strand of duplex siRNA is incorporated into RNA-induced silenc-ing complex (RISC) protein. Subsequently, the incorporated siRNA stand guides RISC to its homologous target mRNA for endonucleolytic cleavage.
Potent knockdown of the target gene with high sequence specificity makes siRNA a promising therapeutic strategy. Three different ways are commonly used for producing siRNA: chemical synthesis, administration of plasmid DNA, and viral vectors encoding small hairpin RNA (shRNA) expression cassette. The transcription of genetic sequence in the plasmid DNA or viral vector leads to the production of an mRNA that has internal self-complementarity, which leads to the formation of double strand with a closed hairpin-like loop at one end (shRNA). The shRNA becomes a sub-strate for the Dicer, leading to endogenous formation of siRNA.
Related Topics
