Bootstrap Percentile Method Confidence Intervals
| Home | | Advanced Mathematics |Chapter: Biostatistics for the Health Sciences: Estimating Population Means
Now that you have learned the bootstrap principle, it is relatively simple to generate percentile method confidence intervals for the mean.
BOOTSTRAP PERCENTILE METHOD CONFIDENCE INTERVALS
Now that you have learned the bootstrap principle,
it is relatively simple to generate percentile method confidence intervals for
the mean. The advantages of the bootstrap confidence interval are that (1) it
does not rely on any parametric distributional assumptions; (2) there is no
reliance on a central limit theorem; and (3) there are no complicated formulas
to memorize. All you need to know is the boot-strap principle. Suppose we have a
random sample of size 10. Consider the pig blood loss data (treatment group)
shown in Table 8.2, which reproduces the treatment data from Table 8.1.
TABLE 8.2. Pig Blood Loss Data (ml)
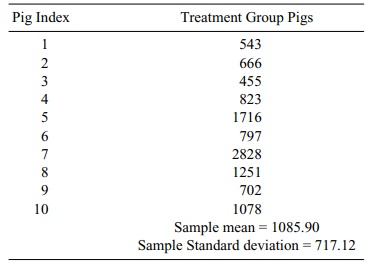
Let us use the method in Section 8.4 based on the t statistic to generate a para-metric
95% confidence interval for the mean. Then we will show you how to gener-ate a
bootstrap percentile method confidence interval based on just 20 bootstrap
samples. We will then show you a better approximation based on 10,000 bootstrap
samples. The result based on 10,000 bootstrap samples requires intensive
comput-ing, which we do using the software package Resampling Stats.
Recall that the parametric confidence interval
based on t is [![]() – C(S/√n), X + C(S/√n)], where S is the sample standard deviation,
– C(S/√n), X + C(S/√n)], where S is the sample standard deviation, ![]() is the sample mean, and
C is the constant taken from the t distribution with n – 1 degrees of freedom, where n
is the sample size and C satisfies
the relationship P(–C ≤ t ≤ C) =
0.95. In this case, n = 10 and df = n
– 1 = 9. From the table of Student’s t
we see that C = 2.2622.
is the sample mean, and
C is the constant taken from the t distribution with n – 1 degrees of freedom, where n
is the sample size and C satisfies
the relationship P(–C ≤ t ≤ C) =
0.95. In this case, n = 10 and df = n
– 1 = 9. From the table of Student’s t
we see that C = 2.2622.
Now, in our example, ![]() = 1085.90 ml and s =
717.12 ml. So the confidence in-terval is [1085.9 – 2.2622(717.12/√10, 1085.9 + 2.2622(717.12/√10] =
[1085.9 – 513.01, 1085.9 + 513.01] = [572.89, 1598.91]. Similarly, for a 90%
interval the val-ue for C is 1.8331;
hence, the 90% interval is [1085.9 – 415.7, 1085.9 + 415.7] = [670.2, 1501.6].
= 1085.90 ml and s =
717.12 ml. So the confidence in-terval is [1085.9 – 2.2622(717.12/√10, 1085.9 + 2.2622(717.12/√10] =
[1085.9 – 513.01, 1085.9 + 513.01] = [572.89, 1598.91]. Similarly, for a 90%
interval the val-ue for C is 1.8331;
hence, the 90% interval is [1085.9 – 415.7, 1085.9 + 415.7] = [670.2, 1501.6].
Now let us generate 20 bootstrap samples of size 10
and calculate the mean of each bootstrap sample. We first list the samples
based on their pig index and then we will compute the bootstrap sample values
and estimates. To generate 20 boot-strap samples of size 10 we need 200 uniform
random numbers. The following 10 × 20 table (Table 8.3) provides the 200
uniform random numbers. Each row repre-sents a bootstrap sample. The pig
indices are obtained as follows:
If the uniform random number U is in [0.0, 0.1), the pig index I is 1.
If the uniform random number U is in [0.1, 0.2), the pig index I is 2.
If the uniform random number U is in [0.2, 0.3), the pig index I is 3.
If the uniform random number U is in [0.3, 0.4), the pig index I is 4.
If the uniform random number U is in [0.4, 0.5), the pig index I is 5.
If the uniform random number U is in [0.5, 0.6), the pig index I is 6.
If the uniform random number U is in [0.6, 0.7), the pig index I is 7.
If the uniform random number U is in [0.7, 0.8), the pig index I is 8.
If the uniform random number U is in [0.8, 0.9), the pig index I is 9.
If the uniform random number U is in [0.9, 1.0), the pig index I is 10.
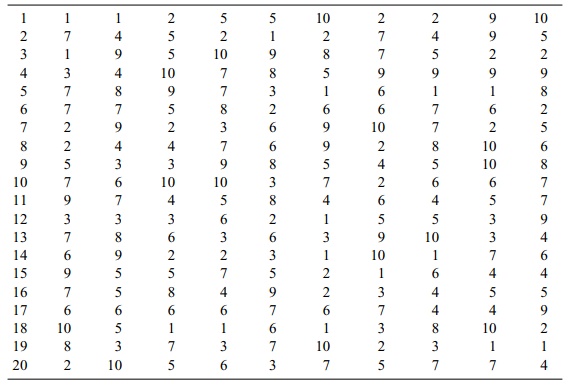
TABLE 8.3. Bootstrap Sample Uniform Random Numbers

In Table 8.4, the indices replace the random
numbers from Table 8.3. Then in Table 8.5, the treatment group values from Table
8.2 replace the indices. The rows in Table 8.5 show the bootstrap sample
averages with the bottom row showing the av-erage of the 20 bootstrap samples.
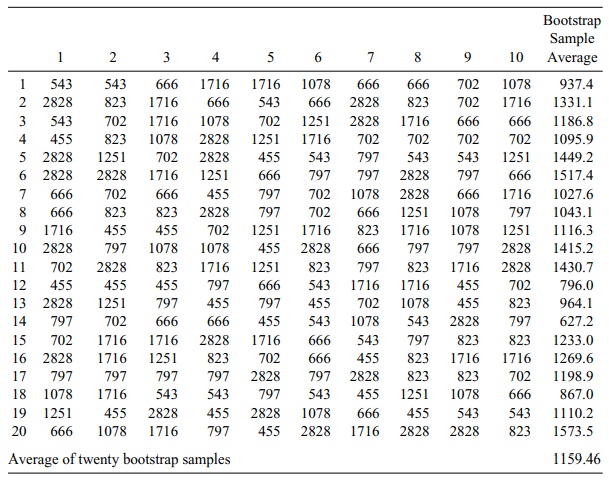
Note in Table 8.5 the similarity of the overall
bootstrap estimates to the sample estimates. For the original sample the
sample, mean was 1085.9 and the estimate of its standard error was 226.77. By
comparison, the bootstrap estimate of the mean is 1159.46 and its bootstrap
estimated standard error is 251.25. The standard error is obtained by computing
a sample standard deviation for the 20 bootstrap sample es-timates in Table
8.4.
Bootstrap percentile confidence intervals are obtained by ordering the bootstrap estimates from smallest to largest. For an approximate 90% confidence interval, the 5th percentile and the 95th percentile are taken as the endpoints of the interval.
TABLE 8.4. Random Pig Indices Based on Table 8.3
Because there are 20 estimates, the interval is from the second smallest to the next to largest, as 5% of the observations are
below the second smallest (1/20) and 5% are above the second largest (1/20).
Consequently, the 90% bootstrap percentile method confidence interval for the
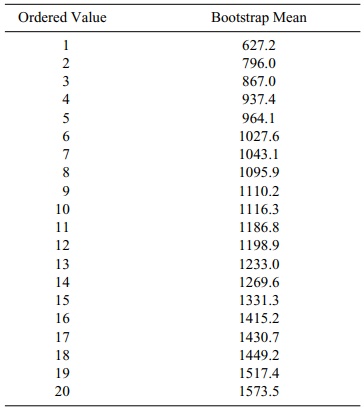
mean is obtained by inspecting Table 8.6, which orders the bootstrap mean
estimates.
Since observation number 2 in increasing rank order
is 796.0 and observation 19 in rank order is 1517.4, the confidence interval is
[796.0, 1517.4]. Compare this to the parametric 90% interval of [670.2,
1501.6]. This difference between the two calculations could be due to the
nonnormality of the data.
We will revisit the results for a random sample of
200 after computing the more precise estimates based on 10,000 bootstrap
samples. Using 10,000 bootstrap sam-ples, we will also be able to compute and
compare the 95% confidence intervals. These procedures will require the use of
the computer program Resampling Stats.
Resampling Stats is a product of the company of the
same name founded by Ju-lian Simon and Peter Bruce to provide software tools to
teach and perform statisti-cal calculations by bootstrap and other resampling
methods. Their software is dis-cussed further in Chapter 16.
Using the Resampling Stats software, we created the
following program (dis-played in italics) in the Resampling Stats language:
data (543 666 455 823 1716 797 2828 1251 702 1078) bdloss
maximize z 15000
mean bdloss mb
stdev bdloss sigb
TABLE 8.5. Bootstrap Sample Blood Loss Values and Averages Based on Pig Indices from Table 8.4
print mb sigb
repeat 10000
sample 10 bdloss bootb
mean bootb mbs$
stdev bootb sigbs$
score mbs$ z
end
histogram z
percentile z (2.5 97.5) k
print mb k
The first line of the code is the data statement. An array is a collection or vector of values stored under a common name and indexed from 1 to n, where n is the ar-ray size. It takes the 10 blood loss values for the pigs and stores it in an array called bdloss; bdloss is an array of size n = 10.
TABLE 8.6. Bootstrap Estimates of Mean Blood Loss in Increasing Order
The next line is the maxsize statement. This statement specifies an array size of 15,000 for the array z. By default, arrays are normally limited to be 1000 in length. So
the n = 15,000 for the array z. We will be able to generate up to
15,000 bootstrap samples (i.e., B =
10,000 for the number of bootstrap samples in this application, but the number
could have been as large as 15,000).
The next two statements, mean and stdev, compute
the sample mean and sample standard deviation, respectively, for the data in
the bdloss array. The results are
stored in the variables mb and sigb for the mean and standard
deviation, respective-ly. The print
statement tells the computer to print out the results.
The repeat
statement then tells the computer how many times to repeat the next several
statements. It starts a loop (like a do
loop in Fortran). The sample
statement tells the computer how to generate the bootstrap samples. The number 10 tells it to sample with replacement
10 times.
The array bdloss
appears in the position to tell the computer to sample from the data in the bdloss array. Then the name bootb is the array to store the
bootstrap sam-ple. The next two statements produce the sample means and
standard deviations for the bootstrap samples. The score statement tells the computer to keep the results for the
means in a vector called z. The end statement indicates the end of the
loop that does the calculations for each of the 10,000 bootstrap samples.
The histogram
statement then takes the results in z
and creates a histogram, auto-matically choosing the number of bins (i.e.,
intervals for the histogram), the bin width and the center of each bin. The percentile statement tells the computer
to list the specified set of percentiles from the distribution determined by
the array of bootstrap means that are stored in z (like the last column in Table 8.5 from the sam-ple of 20
bootstrap estimates of mean blood loss).
When we choose 2.5 and 97.5, these values will
represent the endpoint of a boot-strap percentile method confidence interval at
the 95% confidence level for the mean based on 10,000 bootstrap samples. The
final print statement prints the
sam-ple mean of the original sample and the endpoints of the bootstrap
confidence inter-val. In real time, the program took 1.5 seconds to execute;
the results (in bold face) appeared exactly as follows:
MB = 1085.9
SIGB = 717.12
Vector no. 1: Z
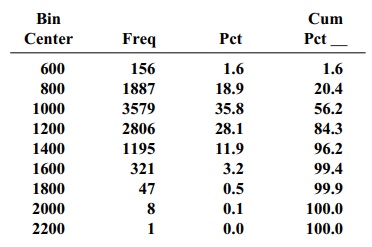
Note: Each bin covers all values within 100 of its center.
MB = 1085.9
K = 727.1 1558.9
Interpreting the output, MB represents the sample mean for the original data and SIGB the standard deviation for the
original data. The histogram is for
Vector no. 1, the array Z of
bootstrap sample means. K is an
array of size n = 2 with its first
el-ement the 2.5 percentile from the histogram of bootstrap means and the
second ele-ment the 97.5 percentile from that histogram.
Using 10,000 random samples, the bootstrap
percentile method 95% confidence interval is [727.1, 1558.9]. Notice that this
is much different from the confidence interval we obtained by assuming a normal
distribution. Recall that that interval was [572.89, 1598.91], which is much
wider than the interval produced by the boot-strap percentile method. This
result is due to the fact that the distribution for the in
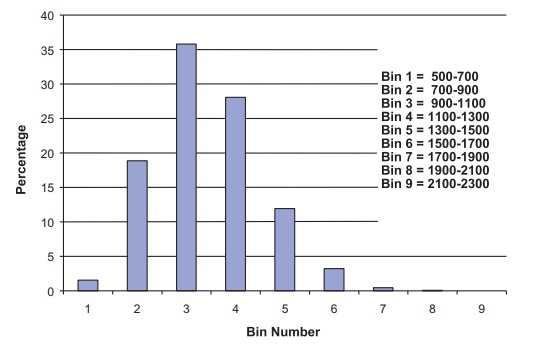
Figure 8.3. Histogram of bootstrap means for the pig treatment group blood loss used for 95% bootstrap percentile method confidence interval.
Not only does the bootstrap give a tighter interval
than the normal approxima-tion, but also the resulting interval is more
realistic based on the sample we ob-served! Figure 8.3 shows the bootstrap histogram
that indicates a skewed distribu-tion for the sampling distribution of the
mean.
To obtain a 90% bootstrap confidence interval using
Resampling Stats, we need only change the percentile statement above to the
following:
percentile z (5.0 95.0) k
The resulting interval is [727.1, 1558.9]. Recall
that, based on only 20 bootstrap samples, we found [796.0, 1517.4] and from
normal distribution theory [670.2, 1501.6]. Again, the two bootstrap results
are not only different from the results ob-tained by using the normal
distribution, but also are more realistic. We see that 20 samples do not yield
an adequate bootstrap interval estimate.
There is a large difference between 20 bootstrap
samples and 10,000 bootstrap samples. The histogram from the Monte Carlo approximation
provides a good ap-proximation to the bootstrap distribution only as the number
of Monte Carlo itera-tions (B)
becomes large. For B as high as
10,000, this distribution and the resulting confidence interval will not change
much if we continue to increase B.
However, when B
is only 20 this result will not be the case. We chose a small value of 20 for B so that we could demonstrate all the
steps of the bootstrap interval estimate without having to resort to the
computer. But to produce an accurate inter
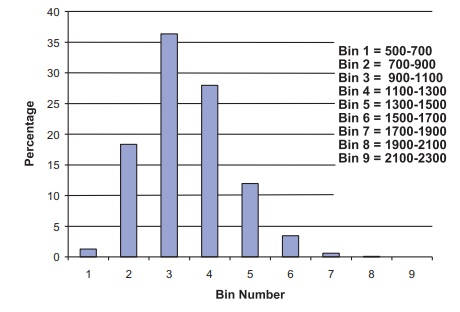
Figure 8.4. Second histogram of bootstrap means for the pig treatment group blood loss. Used for 90% bootstrap percentile method confidence interval.
Subsequently, we found an estimate for the 90%
bootstrap confidence interval by using a different set of 10,000 bootstrap
samples; hence, the histogram (refer to Figure 8.4) is slightly different from
that produced for the 95% confidence interval. The results for this Monte Carlo
approximation are as follows (shown in bold face type):
MB = 1085.9
SIGB = 717.12
Vector no. 1: Z
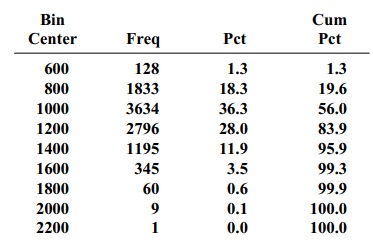
Note: Each bin covers all values within 100 of its center.
Related Topics
