Protein characterization
| Home | | Pharmaceutical Drugs and Dosage | | Pharmaceutical Industrial Management |Chapter: Pharmaceutical Drugs and Dosage: Protein and peptide drug delivery
Biophysical characterization of Protein , Physicochemical characterization of Protein
Protein
characterization
Biophysical
characterization of
Therapeutic
applications of proteins require an understanding of fully elu-cidated
structure, pharmacology, and mechanism of action. In addition, protein behavior
in solution and the impact of chemical properties and components of solutions
on the physical properties of solutions (termed bio-physical characterization)
need to be well defined. Biophysical character-ization of proteins includes the
determination of size, shape, and solution properties of proteins through direct and indirect techniques that
include the following:
· Hydrodynamic protein size measurement by analytical
ultracentri-fugation, gel filtration chromatography, gel electrophoresis,
and/or viscometry.
· Thermodynamic methods such as microcalorimetry and surface
plasma resonance can help delineate the state of protein association and
interactions with other molecules in solution.
·
Particulate formation by protein self-association or
interaction with other components in solution by dynamic light scattering
(DLS).
·
Spectroscopic methods such as circular dichroism (CD) and
thermal melt fluorescence spectroscopy can help determine the stability of
protein conformation in solution.
Physicochemical characterization of Protein
Solubility
Under
physiological conditions, solubility of proteins can vary from the very soluble
to the virtually insoluble. Water solubility of a protein requires
interactions, such as hydrogen bonding and electrostatic inter-actions, of
protein surface with the aqueous medium. The hydrophilic interactions, which
are stronger and predominant in aqueous conditions, are enhanced by the
ionization of functional groups on proteins such as amines and carboxylates.
Ionization of these functional groups is pH dependent. Thus, the solubility of
proteins and peptides is dependent on the pH of the solution.
The
overall charge on a protein can be either positive or negative, depend-ing on
the ionization status of all of its functional groups. A protein is usu-ally
positively charged at a low pH and negatively charged at a high pH. Protein
solubility increases as the pH of the solution moves away from the isoelectric point (IEP) (Figure 25.7), which is the pH at which the mol-ecule
is ionized but has a net zero charge and does not migrate in an elec-tric field
(determined by gel electrophoresis). The presence of both positive and negative
charges on the protein at its IEP leads to a greater tendency for
self-association. As the net charge on the protein changes in any one direction (positive or negative) with a
change in solution pH, the affin-ity of the protein for the aqueous environment
increases and the protein molecules also exert greater electrostatic repulsion
among each other, thus
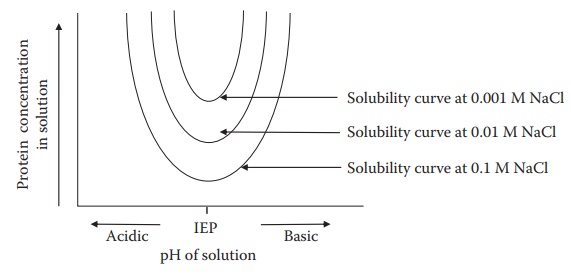
Figure 25.7 A typical profile of protein solubility in solution as a function of
solution pH and salt concentration. IEP, isoelectric point.
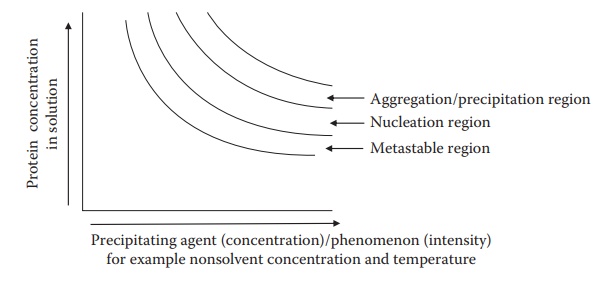
Figure 25.8 Phase behavior of proteins in solution formulation. Typical phases of
physical instability of protein in solution with the addition of a
precipitating agent (such as salt) or change of a precipitation inducing
phenomenon (such as temperature).
The
phase behavior of protein solutions,
that is, whether protein solu-tion is a single-phase solution or has protein
separation (two phases—solid and liquid), is affected by pH, ionic strength,
and temperature (Figure 25.8). Generally,
protein solubility decreases with increasing ionic strength of salts, such as
NaCl and KCl (Figure 25.7). This phenomenon is
called the salting out effect. This phenomenon is used to
concentrate dilute solutions of proteins and
to separate a mixture of proteins (if one of the proteins salts out at a
lower salt concentration than the other). The added salt can then be removed by
dialysis.
Organic
solvents tend to decrease the solubility of proteins by lower-ing the
dielectric constant of the solution (Figure 25.8).
The presence of other highly water-soluble polymers in the solution (cosolutes)
also tends to reduce protein solubility by their interactions with solvent
molecules, thus tying up the solvent and reducing protein–solvent interactions.
This phenomenon is known as the volume
exclusion effect.
Hydrophobicity
Different
amino acids have different degrees of hydrophobicity (Figure
25.4). Overall hydrophobicity or hydrophilicity of a protein is determined
by the nature of functional groups exposed on the surface of the protein. These
are the groups that contribute to protein–solvent and protein–protein
interactions.
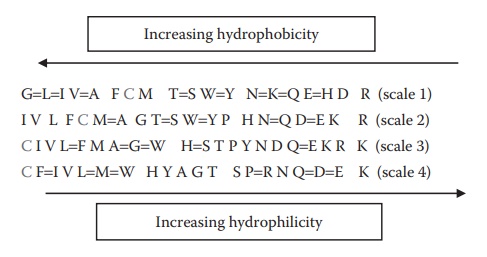

Figure 25.4 Relative hydrophobicity of different amino acids estimated based on either their side-chain sequence (scales 1 and 2) or their typical location in a globular protein structure (scales 3 and 4).
In
an aqueous solution, hydrophobic regions of a polypeptide tend to point away
from the hydrophilic aqueous environment to achieve the ther-modynamically
least energy state of greatest stability. In doing so, the hydrophobic surfaces
of a protein tend to cluster together on the inside of the protein and form
multiple weak van der Waals interactions. These multiple simultaneous weak
hydrophobic interactions are the single most important stabilizing influence of
protein native structure, which also pro-vide flexibility of protein
conformation depending on its solution environ-ment. Thus, in addition to the
stabilizing interactions with the solvent on the surface, including
electrostatic, van der Waals, hydrogen bonds, and ionic interactions,
hydrophobic interactions within and among a protein’s polypeptide chains
stabilize native protein native structure. For example, if alternating
hydrophilic and hydrophobic amino acid sequences in synthetic peptides are at
the optimum distances in space, the molecules coil with the hydrophobic amino
acids on the inside of each coil and the hydrophilic ones to the outside. Thus,
secondary, tertiary, and quaternary structures of polypeptide chains are
important in determining the net hydrophobic or hydrophilic nature of the
protein.
Related Topics
