Structure of Proteins and peptides
| Home | | Pharmaceutical Drugs and Dosage | | Pharmaceutical Industrial Management |Chapter: Pharmaceutical Drugs and Dosage: Protein and peptide drug delivery
Proteins and peptides consist of simple building blocks called amino acids, which are linked together by peptide bonds.
Structure
Proteins
and peptides consist of simple building blocks called amino acids, which are
linked together by peptide bonds. A peptide bond is formed by the nucleophilic
addition of the primary amine of one amino acid to the electropositive
carboxylate carbon of the other amino acid (Figure
25.1). Two amino acids linked together by a peptide bond form a dipeptide;
three amino acids linked together by a peptide bond form a tripeptide; and so
on. Polypeptides consist of a linear chain of several amino acids. Long chains
of amino acids tend to self-associate and fold into three-dimensional
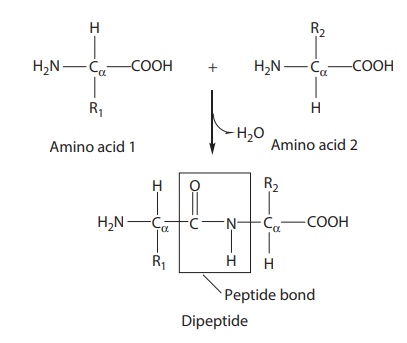
Figure 25.1 Chemical structure of a typical peptide bond. Polypeptides consist of a linear chain of amino acids successively linked via peptide bonds.
As
shown in Figure 25.2, a chain of amino acids
forming a polypeptide through covalent linkages constitutes a protein or
peptide’s primary struc-ture. Spatial
folding of a polypeptide chain through noncovalent interac-tions of neighboring amino acids results in the secondary structure, which consists of
patterns of structural domains such as α-helices and β-sheets. The surfaces of polypeptide
chains, organized into these domains, can further bond with each other through
noncovalent interactions of distant
amino acids, which give the overall structure to one polypeptide chain, called
the tertiary structure. Spatial
interaction of more than one polypep-tide chain to form protein is termed the quaternary structure.
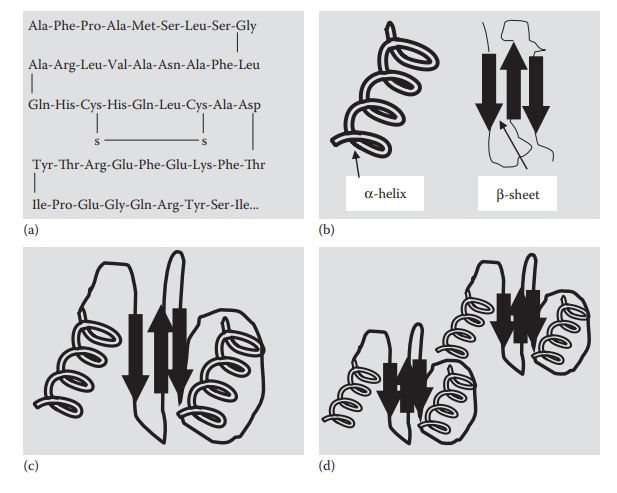
Figure 25.2 Illustration of protein structures: (a) primary structure (amino acid sequence), (b) secondary structure (α-helix and β-sheets), (c) tertiary structure (further folding of the secondary structurally folded protein), and (d) quaternary structure (combination of polypeptides).
Amino acids
There
are 20 naturally occurring amino acids that form the structural basis of all
the proteins and peptides. The chemical structures of these amino acids, along
with their abbreviated and one-letter designations, are pre-sented in Figure 25.3. Each amino acid possesses unique
physicochemical properties governed by their chemical structure.
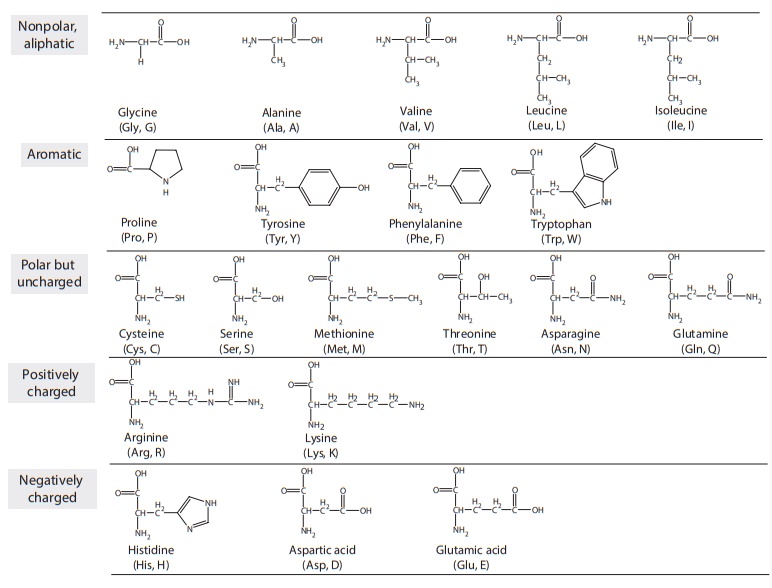
Figure 25.3 Chemical structure of the 20 amino acids commonly found in proteins. The
amino acids may be subdivided into five groups on the basis of side-chain
structure. Their three- and one-letter abbreviations are also listed.
Table 25.2 Hydrophobicity and acidity of amino acids
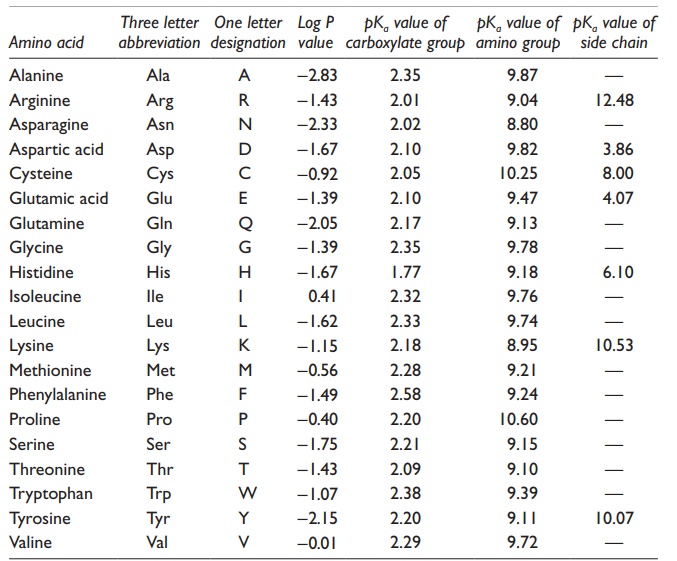
·
Nineteen amino acids contain an amino (–NH2) and
carboxyl (–COOH) group attached to a carbon atom to which various side chains
(–R) are connected. This carbon atom is termed α-carbon because it is next to the
carboxylate group in the structure. The amino acid proline is unusual; in that, its side chain forms a direct covalent
bond with the nitrogen atom of amino group. This is indicated in the higher
hydro-phobic character of proline (higher log P, Table 25.2) compared to most
other amino acids.
·
The α-carbon
has four different groups attached to it and is chiral, except in the case of
glycine. This chirality can lead to two optical isomers, l- and d-amino acids,
which would be mirror images of each other. Natural amino acids are exclusively
l-amino acids.
Amino
acids are classified by the acidity (or basicity) and polarity (or
hydrophilic/hydrophobic nature). The acidity/basicity is indicated by their
ionization constant, the pKa,
whereas the polarity is indicated by log P.
These
constants are defined by the following equations and are listed for each amino
acid in Table 25.2.
An
amino acid backbone can involve the ionization of the acid and/or base:
R-COOH ↔ R-COO− + H+
R-NH2
+ H+ ↔ R-NH3+
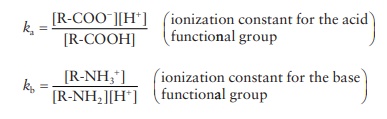

pKa = − log Ka
pKb = − log Kb
p
Ka + pKb =
p K w = 14,
where
p Kw is the ionization
constant of water
and
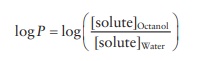
Amino
acids with low pKa values
are acidic, whereas amino acids with high (>7) pK a values (which would correspond to low pKb values) are basic. Amino
acids typically have an acidic carboxylate group and a basic amino group, which
contribute to its acidity or basicity. In addition, the side chain may also
ionize. Thus, there are multiple pKa
values associated with an amino acid. However, in a polypeptide chain, the
carboxylate and amino groups are covalently bonded to neighboring amino acids
(except for the terminal amino acids). In addition, the electron density on the
side chain is influenced by the side chains of other spatially close amino
acids. Thus, ionization constants of amino acids in a protein are different
than those of pure amino acids. Aspartic and glutamic amino acids are
considered acidic because of the presence of ionizable carboxylic acid
functional groups. Arginine, histidine, and lysine contain basic ionizable side
chains and are referred to as basic amino acids.
The
hydrophobic character of amino acids as individual molecules is indicated by
their log P value (Table 25.2). These values are predominantly
influenced by the ionizable carboxylate and amino functional groups. In a
protein structure, these functional groups are covalently bonded.
Hydrophobicity, in the context of protein surface, is primarily influenced
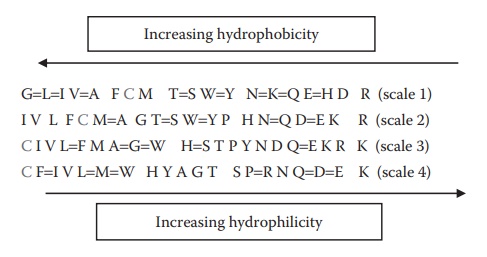

Figure 25.4 Relative hydrophobicity of different amino acids estimated based on either their side-chain sequence (scales 1 and 2) or their typical location in a globular protein structure (scales 3 and 4).
Thus,
the hydrophobic character of amino acids depends on their micro-environment in
the specific protein. In general, the hydrophobic character of an amino acid
has been defined by either (a) physicochemical properties of amino acid side
chains while ignoring the effects of the carboxylate and the amino groups
(scales 1 and 2 in Figure 25.4), or (b) scaling
the prob-ability for an amino acid to be found inside or outside a protein
structure by examining three-dimensional structures of known proteins (scales 3
and 4 in Figure 25.4). The scaling criteria
inherently result in different predic-tions. For example, cysteine typically
forms disulfide bonds in proteins, and stable disulfide bonds are generally
present on the hydrophobic interior of a globular protein. Thus, cysteine is
relatively more hydrophobic by the scaling criterion of its location in a
protein.
Primary structure
The
primary structure of a protein refers to the sequence of amino acids and the
location of disulfide bonds in the constituent polypeptide chain(s) (Figure 25.2). Primary structure determines a
protein’s folding and higher levels of structural organization. However, the
primary structure generally cannot predict the three-dimensional structure and
shape of the proteins in solution.
Secondary structure
Secondary
structure can be described as the local spatial conformation of a polypeptide’s
backbone, excluding the constituent amino acid’s side chains. Common secondary
structural forms are the α-helix
and β-sheets (Figure 25.2). The α-helix results from the helical
coiling of a stretch of hydrophobic amino acids with the hydrophobic groups
facing inside and the hydrophilic groups facing outside the helix. β-sheets, on the other hand, are
characterized by side-by-side hydrogen bonding either within the same chain or
between two different chains, thus exposing the amino acid func-tional groups
to the solvent medium. The chain folding of the secondary structures often
arises from cross-linking through hydrogen bonding or disulfide bridges.
Generally, α-helices are present
in membrane proteins, whereas secreted proteins mostly have β-sheet or irregular structure.
Tertiary structure
Tertiary
structure of a protein refers to the exact three-dimensional struc-ture of its
constituent polypeptide chain(s) (Figure 25.2).
The spatial prox-imity of secondary structural elements determines the tertiary
structure of a polypeptide. Spatially close amino acids on the folded
(secondary structure) polypeptide chains can form attractive hydrogen bond,
ionic, or hydropho-bic interactions, resulting in stabilization of the tertiary
structure. Proteins under physiological conditions assume their distinctive
tertiary structure of minimum free energy, which is a prerequisite for their
biological function.
Quaternary structure
Quaternary
structures are the highest level of protein organization that can be achieved
by proteins that have more than one noncovalently linked constituent
polypeptide chain (Figure 25.2). These
polypeptide chains can associate to form dimers, trimers, and oligomers, which
constitute the qua-ternary structure of a protein. Almost all proteins that are
greater than 100 kDa have a quaternary structure. For example, hemoglobin
consists of nonidentical subunits that associate to form a dimer (heterodimer)
or a tetramer (heterotetramer); glutathione-S-transferase
consists of homotet-ramer (all subunits identical); collagen is a homotrimeric
protein; and the enzyme reverse transcriptase is a heterodimer.
The
stabilization of higher orders of protein structure by multiple weak bonds is
responsible for the flexibility of structure, which is often required for its
functionality. For example, enzymes change conformation upon binding of an
agonist, and membrane ion channels change conformation to facilitate transport
upon ion binding on their surface.
Related Topics
