The Genetic Code
| Home | | Biochemistry |Chapter: Biochemistry : Protein Synthesis
The genetic code is a dictionary that identifies the correspondence between a sequence of nucleotide bases and a sequence of amino acids. Each individual “word” in the code is composed of three nucleotide bases. These genetic words are called codons.
THE GENETIC CODE
The genetic code is a
dictionary that identifies the correspondence between a sequence of nucleotide
bases and a sequence of amino acids. Each individual “word” in the code is
composed of three nucleotide bases. These genetic words are called codons.
A. Codons
Codons are presented in
the mRNA language of adenine (A), guanine (G), cytosine (C), and uracil (U).
Their nucleotide sequences are always written from the 5I -end to the 3I -end.
The four nucleotide bases are used to produce the three-base codons. There are,
therefore, 64 different combinations of bases, taken three at a time (a triplet
code) as shown in the table in Figure 31.2.
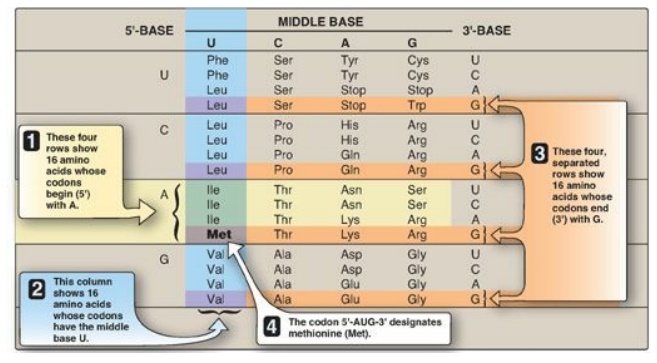
Figure 31.2 Use of the genetic code table to translate the codon AUG. A = adenine; G = guanine; C = cytosine; U = uracil. The abbreviations for many common amino acids are shown as examples.
1. How to translate a codon: This table (or “dictionary”) can
be used to translate any codon and, thus, to determine which amino acids are
coded for by an mRNA sequence. For example, the codon 5I -AUG-3I codes for
methionine ([Met] see Figure 31.2). [Note: AUG is the initiation (start) codon
for translation.] Sixty-one of the 64 codons code for the 20 common amino
acids.
2. Termination (“stop,” or “nonsense”) codons: Three of the codons, UAA, UAG, and UGA, do not code for amino acids but, rather, are termination codons. When one of these codons appears in an mRNA sequence, synthesis of the polypeptide coded for by that mRNA stops.
B. Characteristics of the genetic code
Usage of the genetic
code is remarkably consistent throughout all living organisms. It is assumed
that once the standard genetic code evolved in primitive organisms, any mutation
that altered its meaning would have caused the alteration of most, if not all,
protein sequences, resulting in lethality. Characteristics of the genetic code
include the following.
1. Specificity: The genetic code is specific (unambiguous),
because a particular codon always codes for the same amino acid.
2. Universality: The genetic code is virtually universal insofar as its specificity has been conserved from very early stages of evolution, with only slight differences in the manner in which the code is translated. [Note: An exception occurs in mitochondria, in which a few codons have meanings different than those shown in Figure 31.2. For example, UGA codes for tryptophan (Trp).]
3. Degeneracy: The genetic code is degenerate (sometimes called
redundant). Although each codon corresponds to a single amino acid, a given
amino acid may have more than one triplet coding for it. For example, arginine
(Arg) is specified by six different codons (see Figure 31.2). Only Met and Trp
have just one coding triplet.
4. Nonoverlapping and commaless: The genetic code is nonoverlapping
and commaless, meaning that the code is read from a fixed starting point as a
continuous sequence of bases, taken three at a time without any punctuation
between codons. For example, AGCUGGAUACAU is read as AGC UGG AUA CAU.
C. Consequences of altering the nucleotide sequence
Changing a single
nucleotide base on the mRNA chain (a “point mutation”) can lead to any one of
three results (Figure 31.3).
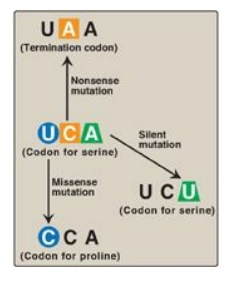
Figure 31.3 Possible effects of changing a single nucleotide base in the coding region of a messenger RNA chain. A = adenine; C = cytosine; U = uracil.
1. Silent mutation: The codon containing the changed
base may code for the same amino acid. For example, if the serine (Ser) codon
UCA is given a different third base, U, to become UCU, it still codes for Ser.
This is termed a “silent” mutation.
2. Missense mutation: The codon containing the changed
base may code for a different amino acid. For example, if the Ser codon UCA is
given a different first base, C, to become CCA, it will code for a different
amino acid (in this case, proline [Pro]). The substitution of an incorrect
amino acid is called a “missense” mutation.
3. Nonsense mutation: The codon containing the changed base may become a termination codon. For example, if the Ser codon UCA is given a different second base, A, to become UAA, the new codon causes termination of translation at that point and the production of a shortened (truncated) protein. The creation of a termination (stop) codon at an inappropriate place is called a “nonsense” mutation.
4. Other mutations: These can alter the amount or
structure of the protein produced by translation.
a. Trinucleotide repeat expansion: Occasionally, a sequence of three
bases that is repeated in tandem will become amplified in number so that too
many copies of the triplet occur. If this happens within the coding region of a
gene, the protein will contain many extra copies of one amino acid. For
example, amplification of the CAG codon leads to the insertion of many extra
glutamine residues in the huntingtin protein, causing the neurodegenerative
disorder Huntington disease (Figure 31.4). The additional glutamines result in
changes in secondary structure that cause the accumulation of protein
aggregates. If the trinucleotide repeat expansion occurs in an untranslated
region (UTR) of a gene, the result can be a decrease in the amount of protein
produced, as seen in fragile X syndrome and myotonic dystrophy. Over 20 triplet
expansion diseases are known. [Note: In fragile X syndrome, the most common
cause of intellectual disability, the expansion results in gene silencing
through DNA hypermethylation.]
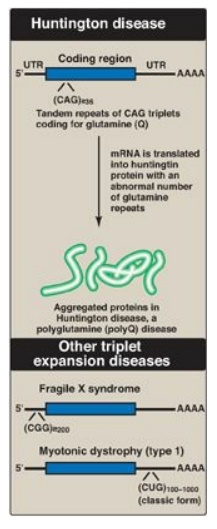
Figure 31.4 Tandem triplet repeats in messenger RNA (mRNA) causing Huntington disease and other triplet expansion diseases. [Note: In unaffected individuals, the number of repeats in the huntingtin protein is fewer than 27, in fragile X mental retardation protein it is 5-44, and in myotonic dystrophy protein kinase it is 5-34.] UTR = untranslated region; A = adenine; C = cytosine; G= guanine; U = uracil; Q = single letter abbreviation for glutamine.
b. Splice-site mutations: Mutations at splice sites can
alter the way in which introns are removed from pre-mRNA molecules, producing
aberrant proteins. [Note: In myotonic dystrophy, a muscle disorder, gene
silencing is the result of splicing alterations due to triplet expansion.]
c. Frame-shift mutations: If one or two nucleotides are
either deleted from or added to the coding region of a mRNA, a frame-shift
mutation occurs, altering the reading frame. This can result in a product with
a radically different amino acid sequence or a truncated product due to the
creation of a termination codon (Figure 31.5). If three nucleotides are added,
a new amino acid is added to the peptide, or, if three are deleted, an amino
acid is lost. Loss of three nucleotides maintains the reading frame but can
result in serious pathology. For example, cystic fibrosis (CF), a chronic,
progressive, inherited disease that primarily affects the pulmonary and
digestive systems, is most commonly caused by deletion of three nucleotides
from the coding region of a gene, resulting in the loss of phenylalanine at the
508th position (DF508) in the protein encoded by that gene. This DF508 mutation
prevents normal folding of the protein, CF transmembrane conductance regulator
(CFTR), leading to its destruction by the proteasome. CFTR normally functions
as a chloride channel in epithelial cells, and its loss results in the
production of thick, sticky secretions in the lungs and pancreas, leading to
lung damage and digestive deficiencies. The incidence of CF is highest (1 in
3300) in those of Northern European origin. In over 70% of individuals with CF,
the DF508 mutation is the cause of the disease.
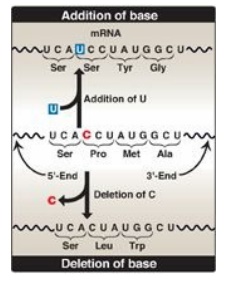
Figure 31.5 Frame-shift
mutations as a result of addition or deletion of a base can cause an alteration
in the reading frame of messenger RNA (mRNA). A = adenine; C = cytosine; G =
guanine; U = uracil.
Related Topics
