Transcription of Prokaryotic Genes
| Home | | Biochemistry |Chapter: Biochemistry : RNA Structure, Synthesis, and Processing
The structure of RNA polymerase (RNA pol), the signals that control transcription, and the varieties of modification that RNA transcripts can undergo differ among organisms, and particularly from prokaryotes to eukaryotes.
TRANSCRIPTION OF PROKARYOTIC GENES
The structure of RNA polymerase (RNA pol), the signals that control transcription, and the varieties of modification that RNA transcripts can undergo differ among organisms, and particularly from prokaryotes to eukaryotes. Therefore, the discussions of prokaryotic and eukaryotic transcription are presented separately.
A. Properties of prokaryotic RNA polymerase
In bacteria, one
species of RNA pol synthesizes all of the RNA except for the short RNA primers
needed for DNA replication [Note: RNA primers are synthesized by a specialized
enzyme, primase.] RNA pol is a multisubunit enzyme that recognizes a nucleotide
sequence (the promoter region) at the beginning of a length of DNA that is to
be transcribed. It next makes a complementary RNA copy of the DNA template
strand, and then recognizes the end of the DNA sequence to be transcribed (the
termination region). RNA is synthesized from its 5I -end to its 3I -end,
antiparallel to its DNA template strand. The template is copied as it is in DNA
synthesis, in which a guanine (G) on the DNA specifies a cytosine (C) in the
RNA, a C specifies a G, a thymine (T) specifies an adenine (A), but an A
specifies a uracil (U) instead of a T (Figure 30.5). The RNA, then, is
complementary to the DNA template (antisense, minus) strand and identical to
the coding (sense, plus) strand, with U replacing T. Within the DNA molecule,
regions of both strands can serve as templates for transcription. For a given
gene, however, only one of the two DNA strands can be the template. Which
strand is used is determined by the location of the promoter for that gene.
Transcription by RNA pol involves a core enzyme and several auxiliary proteins:
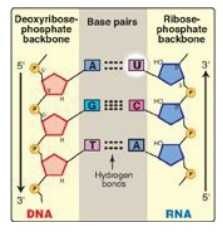
Figure 30.5 Antiparallel,
complementary base pairs between DNA and RNA. T= thymine; A = adenine; C =
cytosine; G = guanine; U = uracil.
1. Core enzyme: Five of the enzyme’s peptide subunits, 2α, 1βI, 1β , and 1Ω, are required for enzyme
assembly (α, Ω) template binding (βI),
and the 5I →3I RNA polymerase activity (β), and are referred to as the core
enzyme (Figure 30.6). However, this enzyme lacks specificity (that is, it
cannot recognize the promoter region on the DNA template).
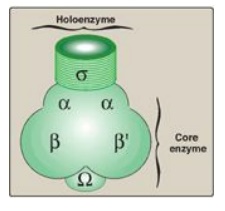
Figure 30.6 Components of prokaryotic RNA polymerase.
2. Holoenzyme: The s subunit (“sigma factor”) enables RNA pol to
recognize promoter regions on the DNA. The s subunit plus the core enzyme make
up the holoenzyme. [Note: Different s factors recognize different groups of
genes.]
B. Steps in RNA synthesis
The process of transcription of a typical gene of Escherichia coli (E. coli) can be divided into three phases: initiation, elongation, and termination. A transcription unit extends from the promoter to the termination region, and the initial product of transcription by RNA pol is termed the primary transcript.
1. Initiation: Transcription begins with the binding of the RNA
pol holoenzyme to a region of the DNA known as the promoter, which is not
transcribed. The prokaryotic promoter contains characteristic consensus
sequences (Figure 30.7). [Note: Consensus sequences are idealized sequences in
which the base shown at each position is the base most frequently (but not
necessarily always) encountered at that position.] Those that are recognized by
prokaryotic RNA polymerase s factors include:
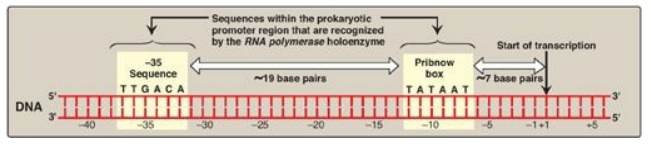
Figure 30.7 Structure of the
prokaryotic promoter region. T = thymine; G = guanine; A = adenine; C =
cytosine.
a. –35 Sequence: A consensus sequence (5-TTGACA-3), centered about
35 bases to the left of the transcription start site (see Figure 30.7), is the
initial point of contact for the holoenzyme, and a closed complex is formed. [Note:
The regulatory sequences that control transcription are, by convention,
designated by the 5→3 nucleotide sequence on the coding strand. A base in the
promoter region is assigned a negative number if it occurs prior to (to the
left of, toward the 5I -end of, or “upstream” of) the transcription start site.
Therefore, the TTGACA sequence is centered at approximately base –35. The first
base at the transcription start site is assigned a position of +1. There is no
base designated “0”.]
b. Pribnow box: The holoenzyme moves and covers a second consensus
sequence (5I -TATAAT-3I ), centered at about –10 (see Figure 30.7), which is
the site of initial DNA melting (unwinding). Melting of a short stretch (about
14 bases) converts the closed complex to an open complex known as a
transcription bubble. [Note: A mutation in either the –10 or the –35 sequence
can affect the transcription of the gene controlled by the mutant promoter.]
2. Elongation: Once the promoter region has been recognized and
bound by the holoenzyme, local unwinding of the DNA helix continues (Figure
30.8), mediated by t h e polymerase. [Note: Unwinding generates supercoils in
the DNA that can be relieved by DNA topoisomerases.] RNA pol begins to
synthesize a transcript of the DNA sequence, and several short pieces of RNA
are made and discarded. The elongation phase is said to begin when the
transcript (typically starting with a purine) exceeds ten nucleotides in
length. Sigma is then released, and the core enzyme is able to leave (“clear”)
the promoter and move along the template strand in a processive manner, serving
as its own sliding clamp. During transcription, a short DNA-RNA
hybrid helix is formed (see Figure 30.8). Like DNA pol, RNA pol uses nucleoside
triphosphates as substrates and releases pyrophosphate each time a nucleoside
monophosphate is added to the growing chain. As with replication, transcription
is always in the 5I →3I direction. In contrast to DNA pol, RNA pol does not
require a primer and does not appear to have 3I →5I exonuclease (proofreading)
activity.
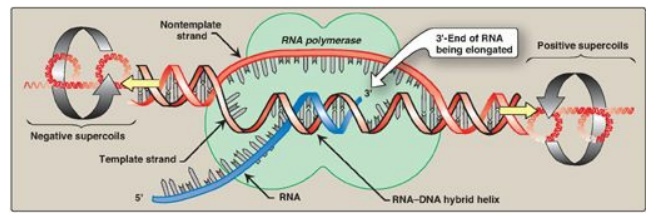
Figure 30.8 Local unwinding of DNA caused by RNA polymerase and formation of an open initiation complex.
3. Termination: The elongation of the single-stranded RNA chain
continues until a termination signal is reached. Termination can be intrinsic
(spontaneous) or dependent upon the participation of a protein known as the ρ
(rho) factor.
a. ρ-Independent termination: Seen with most prokaryotic genes,
this requires that a sequence in the DNA template generates a sequence in the
nascent (newly made) RNA that is self-complementary (Figure 30.9). This allows
the RNA to fold back on itself, forming a GC-rich stem (stabilized by hydrogen
bonds) plus a loop. This structure is known as a “hairpin.” Additionally, just
beyond the hairpin, the RNA transcript contains a string of Us at the 3I -end.
The bonding of these Us to the complementary As of the DNA template is weak.
This facilitates the separation of the newly synthesized RNA from its DNA
template, as the double helix “zips up” behind the RNA polymerase.
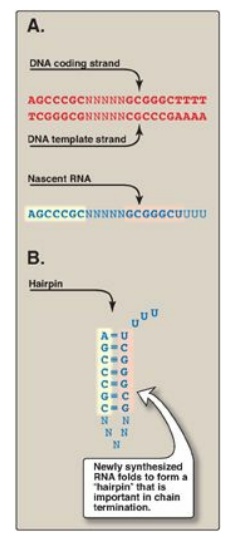
Figure 30.9 Rho-independent termination of prokaryotic transcription. A. DNA template sequence generates a self-complementary sequence in the nascent RNA. B. Hairpin structure formed by the RNA. “N” represents a noncomplementary base; A = adenine, T thymine; G = guanine; C = cytosine; U = uracil. [Note: Termination of eukaryotic transcription is not well understood.]
b. r-Dependent termination: This requires the participation of an additional protein, rho (r), which is a hexameric ATPase with helicase activity. Rho binds a C-rich “rho recognition site” near the 5I -end of the nascent RNA and, using its ATPase activity, moves along the RNA until it reaches the RNA pol paused at the termination site. The ATP-dependent helicase activity of r separates the RNA- DNA hybrid helix, causing the release of the RNA.
4. Action of antibiotics: Some antibiotics prevent bacterial
cell growth by inhibiting RNA synthesis. For example, rifampin (rifampicin)
inhibits transcription by binding to the β subunit of prokaryotic RNA pol, and
preventing chain extension beyond three nucleotides (Figure 30.10). Rifampin is
important in the treatment of tuberculosis. Dactinomycin (known to biochemists
as actinomycin D) was the first antibiotic to find therapeutic application in
tumor chemotherapy. It binds to the DNA template and interferes with the
movement of RNA pol along the DNA.
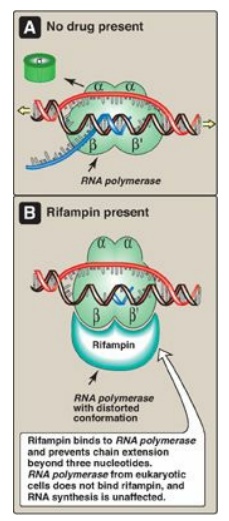
Figure 30.10 Inhibition of
prokaryotic RNA polymerase by rifampin.
Related Topics
