Chapter Summary, Questions Answers - DNA Structure, Replication, and Repair
| Home | | Biochemistry |Chapter: Biochemistry : DNA Structure, Replication, and Repair
DNA is a polymer of deoxyribonucleoside monophosphates covalently linked by 3I →5I -phosphodiester bonds.
CHAPTER SUMMARY
DNA is a polymer of
deoxyribonucleoside monophosphates covalently linked by 3I →5I -phosphodiester
bonds (Figure 29.32). The resulting long, unbranched chain has polarity, with
both a 5I -end and a 3I -end. The sequence of nucleotides is read 5I →3I . DNA
exists as a double-stranded molecule, in which the two chains are paired in an
antiparallel manner, and wind around each other, forming a double helix.
Adenine pairs with thymine, and cytosine pairs with guanine. Each strand of the
double helix serves as a template for constructing a complementary daughter
strand (semiconservative replication). DNA replication occurs in the S phase of
the cell cycle and begins at the origin of replication. As the two strands
unwind and separate, synthesis occurs at two replication forks that move away
from the origin in opposite directions (bidirectionally). Helicase unwinds the
double helix. As the two strands of the double helix are separated, positive
supercoils are produced in the region of DNA ahead of the replication fork and
negative supercoils behind the fork. DNA topoisomerases types I and II remove
supercoils. DNA polymerases (pol) synthesize new DNA strands only in the 5I →3I
direction. Therefore, one of the newly synthesized stretches of nucleotide
chains must grow in the 5I →3I direction toward the replication fork (leading
strand) and one in the 5I →3I direction away from the replication fork (lagging
strand). DNA pols require a primer. The primer for de novo DNA synthesis is a
short stretch of RNA synthesized by primase. The leading strand only needs one
RNA primer, whereas the lagging strand needs many. In E. coli, DNA chain
elongation is catalyzed by DNA pol III, using 5I -deoxyribonucleoside
triphosphates as substrates. The enzyme “proofreads” the newly synthesized DNA,
removing terminal mismatched nucleotides with its 3I →5I exonuclease activity.
RNA primers are removed by DNA pol I, using its 5I →3I exonuclease activity.
This enzyme fills the gaps with DNA, proofreading as it synthesizes. The final
phosphodiester linkage is catalyzed by DNA ligase. There are at least five
high-fidelity eukaryotic DNA polymerases. Pol α is a multisubunit enzyme, one
subunit of which is a primase. Pol α 5I →3I polymerase activity adds a short
piece of DNA to the RNA primer. Pol e completes DNA synthesis on the leading
strand, whereas pol d elongates each lagging strand fragment. Pol β is involved
with DNA repair, and pol γ replicates mitochondrial DNA. Pols e, d, and g u s e
3I →5I exonuclease activity to proofread. Nucleoside analogs containing
modified sugars can be used to block DNA chain growth. They are useful in
anticancer and antiviral chemotherapy. Telomeres are stretches of highly
repetitive DNA complexed with protein that protect the ends of linear
chromosomes. As most cells divide and age, these sequences are shortened,
contributing to senescence. In cells that do not senesce (for example, germline
and cancer cells), telomerase employs its enzyme component reverse
transcriptase to extend the telomeres, using its RNA as a template. There are
five classes of positively charged histone (H) proteins. Two each of histones H2A,
H2B, H3, and H4 form an octameric structural core around which DNA
is wrapped creating a nucleosome. The DNA connecting the nucleosomes, called
linker DNA, is bound to H1. Nucleosomes can be packed more tightly to form a
nucleofilament. Additional levels of organization create a chromosome. Most DNA
damage can be corrected by excision repair involving recognition and removal of
the damage by repair proteins, followed by replacement in E. coli by DNA pol
and joining by ligase. Ultraviolet light can cause thymine dimers that are
recognized and removed by uvrABC proteins of nucleotide excision repair.
Defects in the XP proteins needed for thymine dimer repair in humans result in
xeroderma pigmentosum. Mismatched bases are repaired by a similar process of
recognition and removal by Mut proteins in E. coli. The extent of methylation
is used for strand identification in prokaryotes. Defective mismatch repair by
homologous proteins in humans is associated with hereditary nonpolyposis
colorectal cancer. Abnormal bases (such as uracil) are removed by glycosylases
in base excision repair, and the sugar phosphate at the apyrimidinic or
apurinic (AP) site is cut out. Double-strand breaks in DNA are repaired by
nonhomologous end-joining (error prone) and homologous recombination.
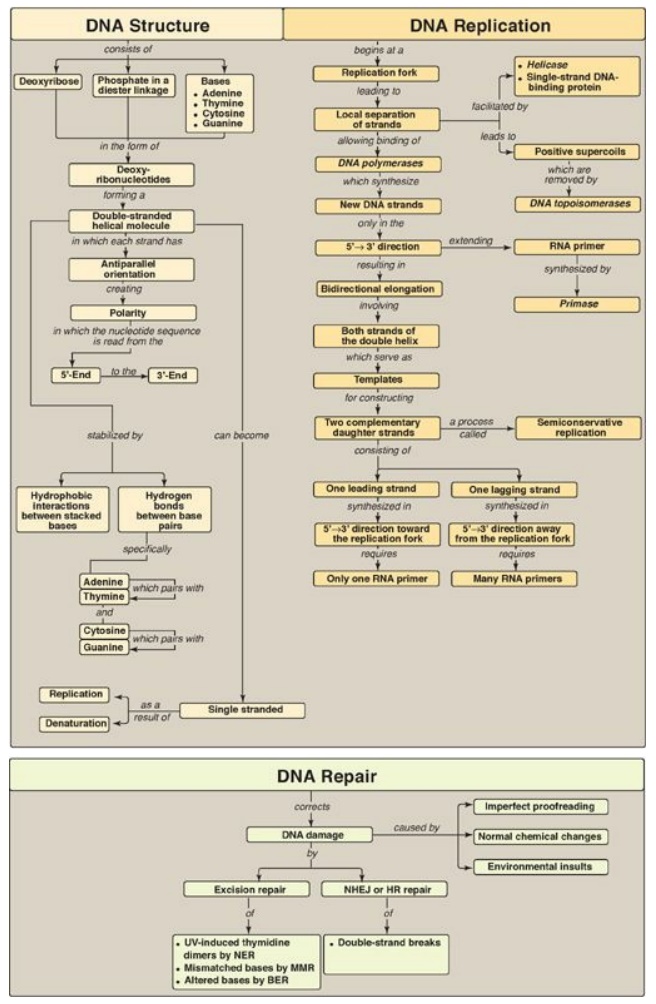
Figure 29.32 Key concept map
for DNA structure, replication, and repair. Key concept map for DNA structure,
replication, and repair. NHEJ = nonhomologous end-joining; HR = homologous recombination;
NER = nucleotide excision repair; MMR = mismatch repair; BER = base excision
repair; UV = ultraviolet light.
Study Questions
Choose the ONE best answer.
29.1 A 10-year-old girl is brought by her parents
to the dermatologist. She has many freckles on her face, neck, arms, and hands,
and the parents report that she is unusually sensitive to sunlight. Two basal
cell carcinomas are identified on her face. Based on the clinical picture,
which of the following processes is most likely to be defective in this
patient?
A. Repair of
double-strand breaks by error-prone homologous recombination
B. Removal of
mismatched bases from the 3I -end of Okazaki fragments by a methyl-directed
process
C. Removal of pyrimidine dimers from DNA by
nucleotide excision repair D. Removal of uracil from DNA by base excision
repair
Correct answer = C. The sensitivity to sunlight,
extensive freckling on parts of the body exposed to the sun, and presence of
skin cancer at a young age indicate that the patient most likely suffers from
xeroderma pigmentosum (XP). These patients are deficient in any one of several
XP proteins required for nucleotide excision repair of pyrimidine dimers in
ultraviolet light–damaged DNA. Double-strand breaks are repaired by
nonhomologous end-joining (error prone) or homologous recombination (error
free). Methylation is not used for strand discrimination in eukaryotic mismatch
repair. Uracil is removed from DNA molecules by a specific glycosylase in base
excision repair, but a defect here does not cause XP.
29.2 Telomeres are complexes of DNA and protein
that protect the ends of linear chromosomes. In most normal human somatic
cells, telomeres shorten with each division. In stem cells and in cancer cells,
however, telomeric length is maintained. In the synthesis of telomeres:
A. telomerase, a ribonucleoprotein, provides both
the RNA and the protein needed for synthesis.
B. the RNA of
telomerase serves as a primer.
C. the RNA of
telomerase is a ribozyme.
D. the protein of telomerase
is a DNA-directed DNA polymerase.
E. the shorter 3I →5I
strand gets extended.
F. the direction of
synthesis is 3I →5I .
Correct answer = A. Telomerase is a ribonucleoprotein
particle required for telomere maintenance. Telomerase contains an RNA that
serves as the template, not the primer, for the synthesis of telomeric DNA by
the reverse transcriptase of telomerase. Telomeric RNA has no catalytic
activity. As a reverse transcriptase, telomerase synthesizes DNA using its RNA
template and so is an RNA-directed DNA polymerase. The direction of synthesis,
as with all DNA synthesis, is 5I →3I , and it is the 3I -end of the already
longer 5I →3I strand that gets extended.
29.3 While studying the structure of a small gene
that was sequenced during the Human Genome Project, an investigator notices
that one strand of the DNA molecule contains 20 As, 25 Gs, 30 Cs, and 22 Ts.
How many of each base is found in the complete double-stranded molecule?
A.A=40,G=50,C=60,T=44
B.A=44,G=60,C=50,T=40
C.A=45,G=45,C=52,T=52
D.A=50,G=47,C=50,T=47
E.A=42,G=55,C=55,T=42
Correct answer = E. The two DNA strands are
complementary to each other, with A base-paired with T and G base-paired with
C. So, for example, the 20 As on the first strand would be paired with 20 Ts on
the second strand, the 25 Gs on the first strand would be paired with 25 Cs on
the second strand, and so forth. When these are all added together, the correct
numbers of each base are indicated in choice E. Notice that, in the correct
answer, A = T and G = C.
29.4 List the order in which the following enzymes
participate in prokaryotic replication.
A. Ligase
B. Polymerase I (3I →5I
exonuclease activity)
C. Polymerase I (5I →3I
exonuclease activity)
D. Polymerase I (5I →3I
polymerase activity)
E. Polymerase III
F. Primase
Correct answer: F, E, C, D, B, A. Primase makes the RNA primer;
Polymerase III extends the primer with DNA (and proofreads); polymerase I
removes the primer with its 5I →3I exonuclease activity, fills in the gap with
its 5I →3I polymerase activity, and removes errors with its 3I →5I exonuclease
activity; and ligase makes the 5I →3I phosphodiester bond that links the DNA
made by polymerase I and polymerase III.
29.5 Dideoxynucleotides lack a 3I -hydroxyl group.
Why would incorporation of a dideoxynucleotide into DNA stop replication?
The lack of the 3I -OH
group prevents formation of the 3I -hydroxyl → 5I -phosphate bond that links
one nucleotide to the next in DNA.
Related Topics
