Comparing Two or More Survival Curves-The Log Rank Test
| Home | | Advanced Mathematics |Chapter: Biostatistics for the Health Sciences: Analysis of Survival Times
To compare two survival curves in a parametric family of distributions such as the negative exponential or the Weibull, we need to test only the hypothesis that the parameters are equal versus the alternative hypothesis that the parameters differ in some way.
COMPARING TWO OR MORE SURVIVAL CURVES-THE LOG RANK TEST
To compare two survival curves in a parametric
family of distributions such as the negative exponential or the Weibull, we
need to test only the hypothesis that the parameters are equal versus the
alternative hypothesis that the parameters differ in some way. We will not go
into the details of such parametric comparisons. Howev-er, nonparametric
procedures look for differences in survival distributions based on the
information in the Kaplan–Meier curves. In this section, we consider specific
nonparametric tests for two or more survival curves.
The log rank test, a nonparametric procedure for
comparing two or more survival functions, is a test of the null hypothesis
that all the survival functions are the same, versus the alternative that at
least one survival function differs from the rest. The idea is to compare the
observed frequency of deaths or failures for each curve in various time
intervals with what would be expected under the null hypothesis that all the
curves are the same. Details can be found in the original paper [see Man-tel
(1966)] or in Lee (1992), pages 109–112.
Now we will describe a simple chi-square test that
is very similar to the log rank test. For the chi-square test, we simply let O1 be the observed number of
deaths in group 1, O2 the
observed number in group 2, O3
the observed number in group 3, and so on until all the groups have been
enumerated.
A chi-square statistic is determined by computing
the expected numbers E1, E2, E3, etc., of deaths in each group. For this calculation
to hold, all the groups need to come
from the same population of survival times. Then, similar to other chi-square
calculations (refer to Chapter 11), the statistic χ2 = (O1
– E1)2/E1 + (O2 – E2)2/E2 + . . . + (Ok – Ek)2/Ek has approximately a
chi-square distribution with k – 1 degrees of freedom when the null
hypothesis is true. We will go through an ex-ample in detail in which k = 2, and the test statistic is then
chi-square with 1 de-gree of freedom under the null hypothesis.
This simple calculation is taken from Lee (1992),
Example 5.2, page 107. Sup-pose that ten female breast cancer patients are
randomized to receive either cyclic administration of cyclophosphamide,
methatrexate, and fluorouracil (CMF), or no additional treatment after a
radical mastectomy. Five patients are randomized to the CMF treatment arm and
five to the control arm.
We are interested in knowing whether time to
relapse (time in remission) is lengthened by the treatment versus the null
hypothesis that the treatment makes no difference. The results at the end of
the trial are as follows: CMF patient remission times in months are 23, 16+,
18+, 20+, and 24+; the control group remission times are 15, 18, 19, 19, and
20. The plus sign (+) indicates that the data were right-cen-sored, e.g., 16+
means right-censored at 16 months. The events without plus signs refer to remission,
1 case for the CMF group and all 5 cases for control group.
Table 15.5 shows the remission times (T), the number of remissions at
remission time (d1), the
number at risk in group 1 (n1t), the number at risk in group 2
(n2t), ex-pected frequency in group 1 (E1), and expected frequency in group 2 (E2). We will use these terms
to compute a chi-square statistic. In order to complete the table, we list the
remission times for the pooled data in ascending order. The remission times
ranged from 15 to 23 months.
At each time t,
the contribution to E1 is dtn1t/(n1t + n2t) and,
similarly, for E2 it is dtn2t/(n1t + n2t). We know that the observed
number of remissions is 1 for group 1 and
5 for group 2. As we see from the first column in the table, the remission
times are at times 15, 18, 19, 20, and 23 with two remissions at 19. As
described previously, the events without the plus signs are the cases in which
the patients relapsed and the time is the time in remission. For the CMF group
we saw that the only such event was at 23 months for one patient. For the
control group we note that five such events oc-curred at times 15, 18, 19, 19,
and 20. So χ2 = (O1
– E1)2/E1 + (O2 – E2)2/E2 = (1 – 3.75)2/4.75
+ (5 – 2.25)2/2.25 = 1.592 + 3.361 = 4.953. From the chi-square distribu-tion
with 1 degree of freedom, we see that this result is statistically significant
at the 0.05 level (0.05 > p >
0.01). Note that with 1 degree of freedom the critical value for p = 0.05 is 3.841 and for p = 0.01 it is 6.635. Since 4.953 lies
between these values we can conclude
that the p-value is between 0.01 and
0.05. Thus, we may conclude that there are significantly shorter remission
times in the control group.
Now let’s consider an example from the treatment of prostate cancer. A proce-dure called cryoablation is used to remove tumors from the prostate gland. Re-searchers assigned each patient to one of three risk groups (i.e., risk of recurrence) based on measures of severity of the disease prior to the procedure. Then, the re-searchers followed the patients for up to eight years.
TABLE 15.5. Computation of Expected Numbers for Chi-square Test
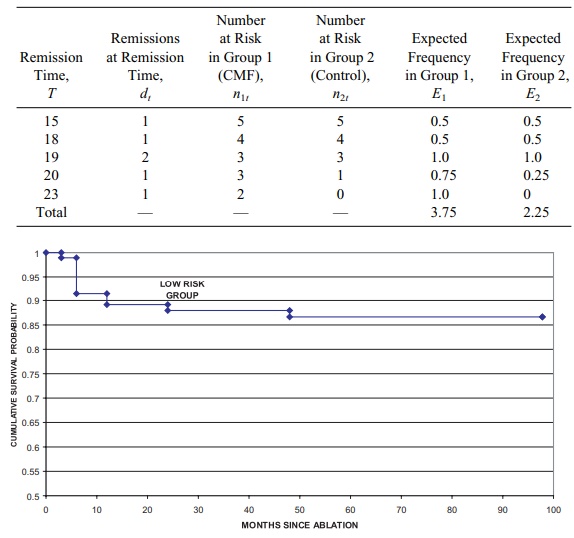
Figure 15.2. Cryoablation biochemical-free survival PSA > 1 criterion.
The three categories of risk were designated as low, moderate, and high. Ka- plan–Meier survival curves were generated for each
risk group; the log rank test was used to compare these survival curves.
Failure was defined as having a prostate-specific antigen lab test result above
1.0 ng/mL. Figures 15.2, 15.3, and 15.4 present the Kaplan–Meier curves.
The curves are very similar. However, the total
sample size was only 561, with 94 patients in the low risk group, 178 in the
medium risk group, and 289 in the high-risk group. The p-value for the log rank test was 0.2597, indicating that the
curves were not statistically significantly different.
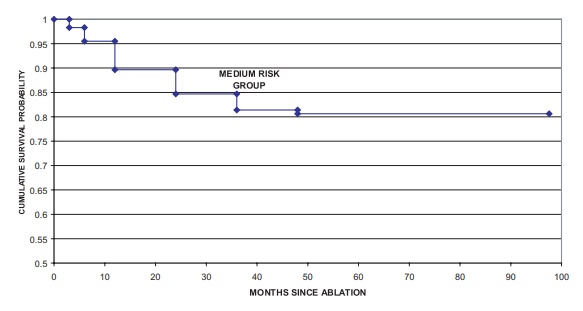
Figure 15.3. Cryoablation biochemical-free
survival PSA > 1 criterion.
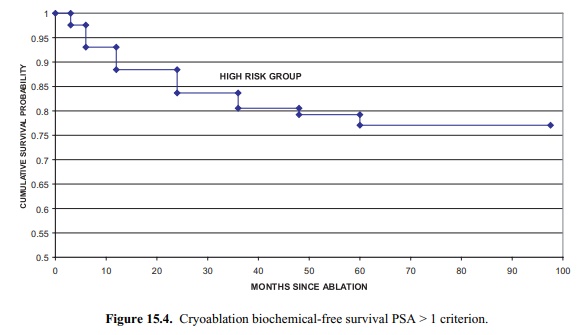
Figure 15.4. Cryoablation biochemical-free
survival PSA > 1 criterion.
With this final example, we conclude Chapter 15.
You have seen that analyses of survival times yield much useful information
regarding the survival of patients and estimation of cure rates. In the next
chapter, we will identify computer software programs that can be used for
survival analyses. Chapter 16 also will present a vari-ety of software packages
that are applicable to many of the statistical techniques covered in this text.
Related Topics
