Overall Nitrogen Metabolism
| Home | | Biochemistry |Chapter: Biochemistry : Amino Acids: Disposal of Nitrogen
Amino acid catabolism is part of the larger process of the metabolism of nitrogen-containing molecules.
OVERALL NITROGEN METABOLISM
Amino acid catabolism
is part of the larger process of the metabolism of nitrogen-containing
molecules. Nitrogen enters the body in a variety of compounds present in food,
the most important being amino acids contained in dietary protein. Nitrogen
leaves the body as urea, ammonia, and other products derived from amino acid
metabolism. The role of body proteins in these transformations involves two
important concepts: the amino acid pool and protein turnover.
A. Amino acid pool
Free amino acids are
present throughout the body, such as in cells, blood, and the extracellular
fluids. For the purpose of this discussion, envision all of these amino acids
as if they belonged to a single entity, called the amino acid pool. This pool
is supplied by three sources: 1) amino acids provided by the degradation of
endogenous (body) proteins, most of which are reutilized; 2) amino acids
derived from exogenous (dietary) protein; and 3) nonessential amino acids
synthesized from simple intermediates of metabolism (Figure 19.2). Conversely,
the amino pool is depleted by three routes: 1) synthesis of body protein; 2)
consumption of amino acids as precursors of essential nitrogen-containing small
molecules; and 3) conversion of amino acids to glucose, glycogen, fatty acids,
and ketone bodies, or oxidation to CO2 + H2O (see Figure
19.2). Although the amino acid pool is small (comprising about 90– 100 g of
amino acids) in comparison with the amount of protein in the body (about 12 kg
in a 70-kg man), it is conceptually at the center of whole-body nitrogen
metabolism.
In healthy, well-fed individuals, the input to the
amino acid pool is balanced by the output. That is, the amount of amino acids
contained in the pool is constant. The amino acid pool is said to be in a
steady state, and the individual is said to be in nitrogen balance.
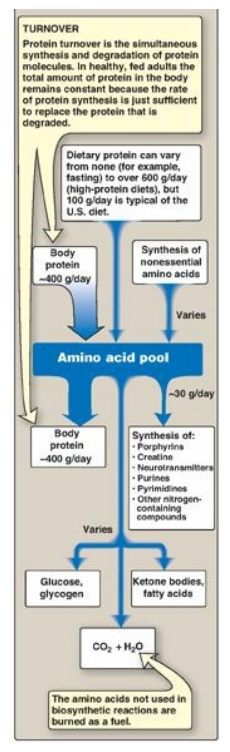
Figure 19.2 Sources and fates
of amino acids.
B. Protein turnover
Most proteins in the
body are constantly being synthesized and then degraded, permitting the removal
of abnormal or unneeded proteins. For many proteins, regulation of synthesis
determines the concentration of protein in the cell, with protein degradation
assuming a minor role. For other proteins, the rate of synthesis is
constitutive (that is, essentially constant), and cellular levels of the
protein are controlled by selective degradation.
1. Rate of turnover: In healthy adults, the total
amount of protein in the body remains constant because the rate of protein
synthesis is just sufficient to replace the protein that is degraded. This
process, called protein turnover, leads to the hydrolysis and resynthesis of
300–400 g of body protein each day. The rate of protein turnover varies widely
for individual proteins. Short-lived proteins (for example, many regulatory
proteins and misfolded proteins) are rapidly degraded, having half-lives
measured in minutes or hours. Long-lived proteins, with half-lives of days to
weeks, constitute the majority of proteins in the cell. Structural proteins,
such as collagen, are metabolically stable and have half-lives measured in
months or years.
2. Protein degradation: There are two major enzyme systems
responsible for degrading proteins: the adenosine triphosphate (ATP)-dependent
ubiquitin-proteasome system of the cytosol, and the ATP-independent degradative
enzyme system of the lysosomes. Proteasomes selectively degrade damaged or
short-lived proteins. Lysosomes use acid hydrolases to nonselectively degrade
intracellular proteins (“autophagy”) and extracellular proteins
(“heterophagy”), such as plasma proteins, that are taken into the cell by
endocytosis.
a. Ubiquitin–proteasome proteolytic pathway: Proteins selected for degradation
by the cytosolic ubiquitin-proteasome system are first modified by the covalent
attachment of ubiquitin (Ub), a small, globular, nonenzymic protein that is
highly conserved across eukaryotic species. Ubiquitination of the target
substrate occurs through isopeptide linkage of the α-carboxyl group of the
C-terminal glycine of Ub to the ε-amino group of a lysine on the protein
substrate by a three-step, enzyme-catalyzed, ATP-dependent process. [Note:
Enzyme 1 (E1, or activating enzyme) activates Ub, which is then transferred to
E2 (conjugating enzyme). E3 (a ligase) identifies the protein to be degraded
and interacts with E2-Ub.] The consecutive addition of four or more Ub
molecules to the target protein generates a polyubiquitin chain. Proteins
tagged with Ub are recognized by a large, barrel-shaped, macromolecular,
proteolytic complex called a proteasome (Figure 19.3). The proteasome unfolds,
deubiquitinates, and cuts the target protein into fragments that are then
further degraded by cytosolic proteases to amino acids, which enter the amino
acid pool. Ub is recycled. It is noteworthy that the selective degradation of
proteins by the ubiquitin-proteosome complex (unlike simple hydrolysis by
proteolytic enzymes) requires energy in the form of ATP.
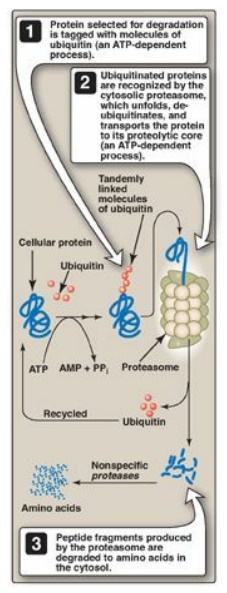
Figure 19.3 The ubiquitin-proteasome degradation pathway of proteins. AMP = adenosine monophosphate; PPi = pyrophosphate.
b. Chemical signals for protein degradation: Because proteins have different half-lives, it is clear that protein degradation cannot be random but, rather, is influenced by some structural aspect of the protein. For example, some proteins that have been chemically altered by oxidation or tagged with ubiquitin are preferentially degraded. The half-life of a protein is also influenced by the amino (N)-terminal residue. For example, proteins that have serine as the N-terminal amino acid are long-lived, with a half-life of more than 20 hours, whereas those with aspartate at their N-terminus have a half-life of only 3 minutes. Additionally, proteins rich in sequences containing proline, glutamate, serine, and threonine (called PEST sequences after the one-letter designations for these amino acids) are rapidly degraded and, therefore, have short half-lives.
Related Topics
