Types of complexes
| Home | | Pharmaceutical Drugs and Dosage | | Pharmaceutical Industrial Management |Chapter: Pharmaceutical Drugs and Dosage: Complexation and protein binding
Depending on the type of interactions involved in complexation, ligand- substrate complexes are classified as follows:
Types
of complexes
Depending
on the type of interactions involved in complexation, ligand– substrate
complexes are classified as follows:
·
Coordination
complexes:
These are covalent complexes that form as
a result of multiple Lewis acid–base reactions in which multiple neutral or
anionic ligands bind a central, cationic substrate through multiple coordinate
covalent bonds. Thus, the ionic covalent bonds are formed when an electron-rich
atom on the ligand bonds with an electropositive atom of or on the substrate by
donating its pair of elec-trons. Tetracycline complexation with divalent heavy
metal cations is an example of a coordination complex.
·
Molecular complexes: These are
noncovalent complexes formed by
multiple attractive interactions between two molecules, such as hydrogen
bonding, electrostatic attraction, van der Waals forces, and hydrophobic
interactions.
Coordination complexes
Metal
complexes are the most common coordination complexes. Their structure involves
one or more central metal atom or cation, surrounded by a number of substrates
with negatively charged ions (such as carboxyl-ate groups) or neutral molecules
possessing lone pair of electrons (such as on nitrogen atoms of amine groups).
The ions or molecules surrounding the metal are called ligands. The number of
bonds formed between the metal ion and the ligand(s) is called the coordination
number of the com-plex. Ligands are generally bound to a metal ion by a
coordinate covalent bond (i.e., donating electrons from a lone electron pair
into an empty metal orbital) and are thus said to be coordinated to the ion.
The
interaction between the metal ion and the ligand is a Lewis acid– base
reaction, in which the ligand (a base) donates a pair of electrons (:) to the
metal ion (an acid) to form the coordinate covalent bond. For example,
Ag+ + 2(: NH3 ) → [Ag(NH3)2 ]+
Where,
silver ion (Ag+) is the central metal ion interacting with ammonia
(NH3) to form the silver–ammonia [Ag(NH 3)2]+
coordination complex. Ligands, such as H2O:, NC−:, and Cl−:
donate a pair of electron in forming a complex. For example, silver–ammonia
complexes can be neutralized with Cl− to form [Ag(NH3)2]Cl.
Several
enzymes involve coordination complexation of their amino acids to one or more
heavy metal atoms. Coordination complexes play a critical role in controlling
the structure and function of many enzymes. Heavy metal ions present in
physiological proteins and enzymes facilitate the for-mation of coordination
complexes that result in the functionality of the protein or the enzyme. For
example, copper ion is present in proteins and enzymes, including hemocyanin,
superoxide dismutase, and cytochrome oxidase. Zinc is present in many proteins
and confers structure and sta-bility, such as crystalline insulin. When present
in deoxyribonucleic acid (DNA)-binding proteins, Zn2+ binds
tetrahedrally with the two histidine and two cysteine residues of the protein
to form a loop (zinc finger), which can fit into the major groove of genomic
double-helical DNA (Figure 6.3).
Several
nonenzymatic molecules of biological significance are coor-dination compounds.
For example, vitamin B12 (cyanocobalamin) is a coordination complex
of cobalt (Figure 6.2), and heme is a
coordination complex of iron with the nitrogens of histidine residues of the
pro-tein (Figure 6.2). Heme proteins of
myoglobin and hemoglobin are iron
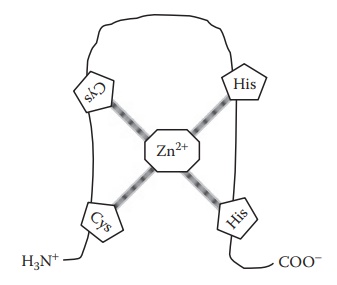
Figure 6.3 Formation of zinc finger due to zinc binding to histidine and cysteine residues in a peptide chain.
Each
heme residue contains one central iron atom in the ferrous oxidation state (Fe2+)
in coordinate bonds with a heterocyclic organic com-pound called porphyrin. The
oxygen carried by heme proteins is bound directly to Fe2+ atom of
the heme group. Oxidation of the iron to the ferric oxidation state (Fe3+)
renders the molecule incapable of binding oxygen.
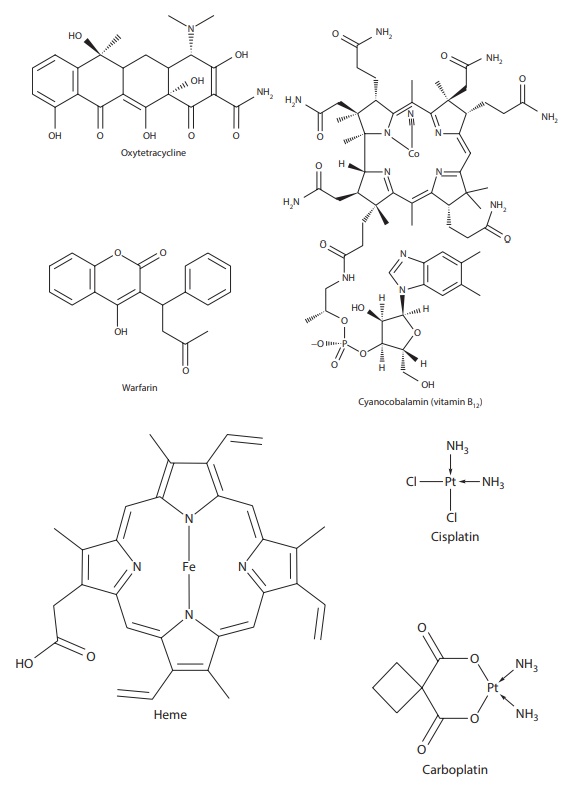
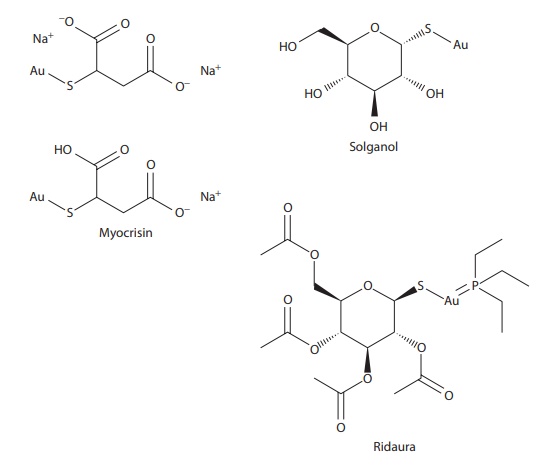
Figure 6.2 Examples of drugs that exist as complexes and/or have a high propensity for forming complexes.
Among drugs, anticancer drugs cisplatin and carboplatin are platinum (II) complexes (Figure 6.2). Rheumatoid arthritis drugs aurothiomalate (Myocrisin®), aurothioglucose (Solganol®), aurothiopropanol sulfonate (Allocrysin®), and nuranofin (Ridaura®) are gold complexes (Figure 6.2).
Molecular complexes
Molecular
complexes involve noncovalent interactions between the ligand and the
substrate, such as electrostatic attraction between oppositely charged ions,
van der Waals forces, hydrogen bonding, and hydrophobic interactions. Molecular
complexes can be subdivided based on the substrate and the ligand involved in
complexation.
1. Small molecule–small molecule complexes
·
Molecules bearing functional groups with opposite polarity
can interact with each other in solution. For example, benzocaine interacts
with caf-feine as a result of a dipole–dipole interaction between the
nucleophilic carbonyl oxygen of benzocaine and the electrophilic nitrogen of
caffeine.
·
Self-association complexes form when drug molecules in
solution inter-act with one another to form dimers, trimers, or higher-order-association
structures, including micelles. For example, daunomycin, mitoxantrone, and
brivanib alaninate are known to self-associate in aqueous solution.
3. Small molecule–large molecule complexes
·
Drugs often interact with macromolecules in vitro. For example, cationically
charged drugs may interact with anionically charged excipients and polymers in
the dosage form, such as tablet, to form a complex. Commonly encountered
anionic hydrophilic polymers in the dosage form include the superdisintegrants
croscarmellose sodium and sodium starch glycolate.
· Drugs can also form complexes with ion-exchange resins. Such
com-plexation can lead to incomplete drug release from the dosage form.
Examples of drugs that can form such complexes include basic drugs amitriptyline,
verapamil, diphenhydramine, alprenolol, and atenolol. Ion-exchange resins that
strongly bind drugs are also used to make sustained-release dosage forms. For
example, ion-exchange resins carboxylic acid and sulfonic acid can bind
cationic drugs and those with quaternary ammonium groups can bind anionic
drugs.
· Several water-soluble pharmaceutical polymers, including
polyethylene glycols (PEGs), polyvinylpyrrolidone (PVP), polystyrene,
carboxymethylcellulose (CMC), and similar polymers containing nucleophilic
oxygen, can form complexes with drugs in solution.
·
Plasma protein binding. Drug–protein complexation between
small-molecule drugs and large protein molecules in the plasma is mediated by
reversible molecular interactions.
·
Enzyme–substrate interactions. Enzyme–substrate interactions
involve very specific noncovalent bonds between various amino acids of the
enzyme folded into the substrate-recognition site. The require-ment of
formation of multiple specific bonds in specific orientation and location within
the substrate-binding site for enzyme action ensures substrate recognition for
activation of the enzyme.
·
Inclusion/occlusion complexes. These complexes involve the
entrapment of one compound in the molecular framework of another. Inclusion
complexes are exemplified by the complexation of hydro-phobic drugs by
cyclodextrin molecules, which totally enclose the substrate. Occlusion
complexes are exemplified by a specific case of cyclodextrin complexes where
only the hydrophobic portion of an amphiphilic molecule is complexed by
cyclodextrin.
Cyclodextrins
are donut-shaped molecules of β-D-glucopyranose
with 6, 7, or 8 cyclic residues of d-glucose, known as α-, β-, or γ-cyclodextrins, respectively (Figure 6.4). The cavity size ranges from 5Å for α-cyclodextrin to 8 Å for γ-cyclodextrin. In addition, several
cyclodextrin derivatives, such as methyl-, dimethyl-, 2-hydroxypropyl, and
sulfobutyl ether substitutions on the hydroxyl groups of the cyclodextrin, lead
to different physicochemi-cal properties. For example, Figure
6.5 shows the ampicillin–cyclodextrin occlusion complex.
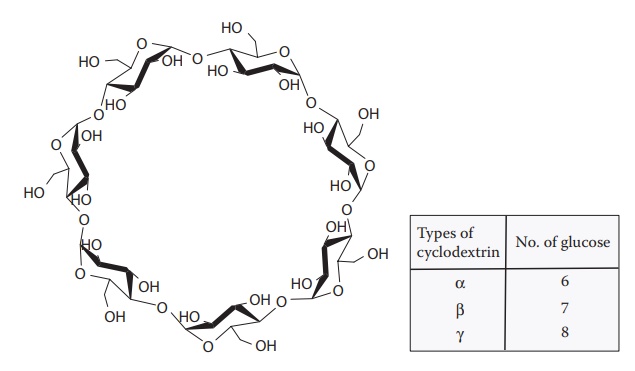
Figure 6.4 Chemical structure of cyclodextrin.
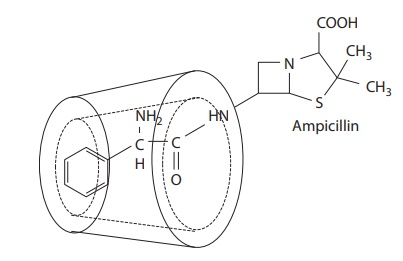
Figure 6.5 Example of complexation of ampicillin by cyclodextrin.
Cyclodextrins
are used to complex hydrophobic molecules or hydro-phobic portions of a
molecule. Complexation is mediated primarily by van der Waals force of
attraction and hydrophobic interaction. The surface of cyclodextrins is highly
hydrophilic because of the multiple hydroxyl (−OH) functional groups that can
hydrogen bond with water. Thus, cyclodex-trins can form reversible
water-soluble inclusion or occlusion complexes of hydrophobic compounds.
Cyclodextrins are nontoxic and do not illicit immune response. Cyclodextrin
complexation can, therefore, serve as an effective means of increasing the
aqueous solubility, stability, absorption, and bioavailability of hydrophobic
drugs. Cyclodextrins have been used to complex and increase the solubility of
various hydrophobic drugs, such as paclitaxel and hydrocortisone.
3. Large molecule–large molecule complexes
·
Large molecule–large molecule complexes are exemplified by
polyacids, which can form hydrogen-bonded complexes with PEGs (Figure 6.6). In addition, PVP can form complexes with
poly(acrylic acids).
·
Base–base interactions in DNA helix through interactions
between the nucleotide bases involve hydrogen bonding and are responsible for
the unique double-helical structure of the double-stranded DNA. The DNA double
helix is stabilized by the hydrogen bond interactions among nucleotides.
Adenine (A) forms two hydrogen bonds with thy-mine (T), and guanine (G) forms
three hydrogen bonds with cytosine (Figure 6.7).
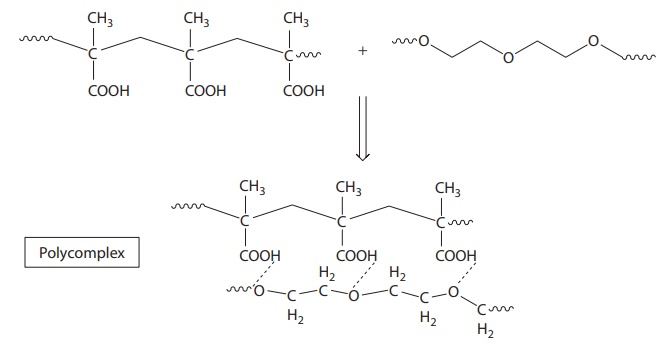
Figure 6.6 Example of
macromolecule–macromolecule interaction. Interaction between polyacid and
polyethylene glycol.
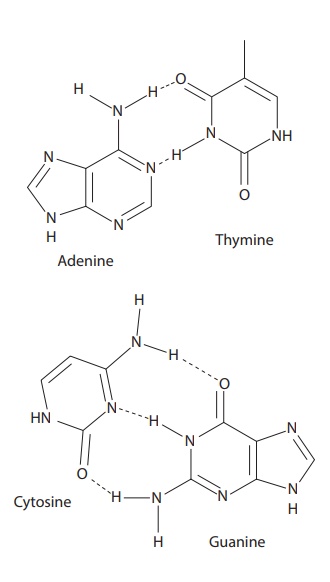
Figure 6.7 Complexation between bases in DNA molecules.
Related Topics
