Analysis of Gene Expression
| Home | | Biochemistry |Chapter: Biochemistry : Biotechnology and Human Disease
The tools of biotechnology not only allow the study of gene structure, but also provide ways of analyzing the mRNA and protein products of gene expression.
ANALYSIS OF GENE EXPRESSION
The tools of
biotechnology not only allow the study of gene structure, but also provide ways
of analyzing the mRNA and protein products of gene expression.
A. Determination of messenger RNA levels
mRNA levels are usually
determined by the hybridization of labeled probes to either mRNA itself or to
cDNA produced from mRNA. [Note: Amplification of cDNA made from mRNA by
retroviral reverse transcriptase (RT) is referred to as RT-PCR.]
1. Northern blots: Northern blots are very similar to
Southern blots (see Figure 33.12), except that the original sample contains a
mixture of mRNA molecules that are separated by electrophoresis, then
transferred to a membrane and hybridized to a radiolabeled probe. The bands
obtained by autoradiography give a measure of the amount and size of particular
mRNA molecules in the sample.
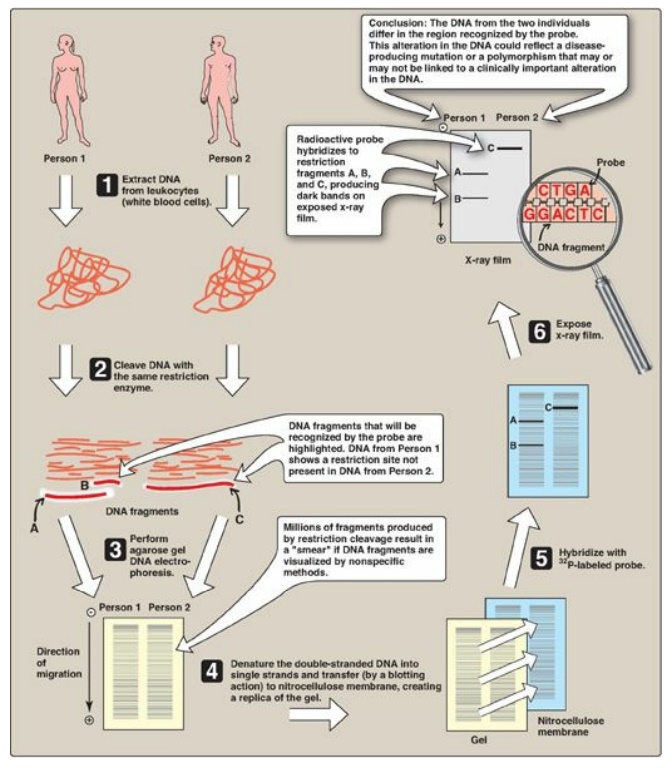
Figure 33.12 Southern blotting procedure. [Note: Nonradiolabeled probes are now commonly used.]
2. Microarrays: DNA microarrays contain thousands of immobilized
ssDNA sequences organized in an area no larger than a microscope slide. These
microarrays are used to analyze a sample for the presence of gene variations or
mutations (genotyping) or to determine the patterns of mRNA production (gene
expression analysis), analyzing thousands of genes at the same time. For
genotyping analysis, the sample is from genomic DNA. For expression analysis,
the population of mRNA molecules from a particular cell type is converted to
cDNA and labeled with a fluorescent tag (Figure 33.22). This mixture is then
exposed to a gene (DNA) chip, which is a glass slide or membrane containing
thousands of tiny spots of DNA, each corresponding to a different gene. The
amount of fluorescence bound to each spot is a measure of the amount of that
particular mRNA in the sample. DNA microarrays are used to determine the
differing patterns of gene expression in two different types of cell (for
example, normal and cancer cells; see Figure 33.22). They can also be used to
subclassify cancers, such as breast cancer, to optimize treatment. [Note:
Microarrays involving proteins and the antibodies or other proteins that
recognize them are being used to identify biomarkers to aid in the diagnosis,
prognosis, and treatment of disease based on a patient’s protein expression
profile. Protein (and DNA) microarrays are important tools in the development
of personalized medicine.]
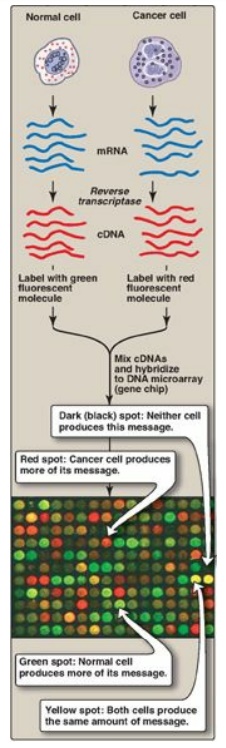
Figure 33.22 Microarray
analysis of gene expression using DNA (gene) chips. [Note: Protein chips are
also used.] mRNA = messenger RNA; cDNA = complementary DNA.
B. Analysis of proteins
The kinds and amounts
of proteins in cells do not always directly correspond to the amounts of mRNA
present. Some mRNAs are translated more efficiently than others, and some
proteins undergo posttranslational modification. When analyzing the abundance
and interactions of a large number of proteins, automated methods involving a
variety of techniques, such as mass spectrometry and two-dimensional
electrophoresis, are used. When investigating one, or a limited number of
proteins, labeled antibodies are used to detect and quantify specific proteins
and to determine posttranslational modifications.
1. Enzyme-linked immunosorbent assays (ELISAs): These assays are performed in the
wells of a plastic microtiter dish. The antigen (protein) is bound to the
plastic of the dish. The probe used consists of an antibody specific for the
particular protein to be measured. The antibody is covalently bound to an
enzyme, which will produce a colored product when exposed to its substrate. The
amount of color produced is proportional to the amount of antibody present and,
indirectly, to the amount of protein in a test sample.
2. Western blots: Western blots (also called
immunoblots) are similar to Southern blots, except that protein molecules in
the sample are separated by electrophoresis and blotted (transferred) to a
membrane. The probe is a labeled antibody, which produces a band at the
location of its antigen.
3. Detecting exposure to human immunodeficiency
virus (HIV): ELISA
and Western blots are commonly used to detect exposure to HIV by measuring the
amount of anti-HIV antibodies present in a patient’s blood sample. ELISAs are
used as the primary screening tool, because they are very sensitive. Because
these assays sometimes give false positives, however, Western blots, which are
more specific, are often used as a confirmatory test (Figure 33.23). [Note:
ELISA and Western blots can only detect HIV exposure after anti-HIV antibodies
appear in the bloodstream. PCR-based testing for HIV is more useful in the
first few months after exposure.]
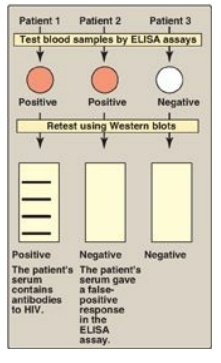
Figure 33.23 Testing for HIV exposure by enzymelinked immunosorbent assays (ELISAs) and Western blots.
C. Proteomics
The study of the
proteome or all the proteins expressed by a genome, including their relative
abundance, distribution, posttranslational modifications, functions, and
interactions with other macromolecules, is known as proteomics. The 20,000 to
25,000 protein-coding genes of the human genome translate into well over 100,000
proteins when posttranscriptional and posttranslational modifications are
considered. Although a genome remains essentially unchanged, the amounts and
types of proteins in any particular cell change dramatically as genes are
turned on and off. [Note: Proteomics (and genomics) required the parallel
development of bioinformatics, the computer-based organization, storage, and
analysis of biologic data.] Figure 33.24 compares some of the analytic
techniques discussed in this chapter.
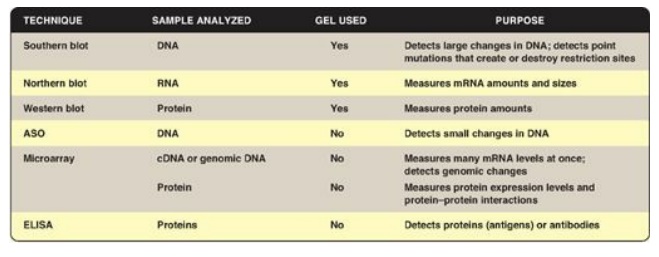
Figure 33.24 Techniques used to
analyze DNA, RNA, and proteins. ASO = allele-specific oligonucleotides. ELISA =
enzymelinked immunosorbent assay; cDNA = complementary DNA; mRNA = messenger
RNA.
Related Topics
