Probes
| Home | | Biochemistry |Chapter: Biochemistry : Biotechnology and Human Disease
Cleavage of large DNA molecules by restriction enzymes produces a bewildering array of fragments.
PROBES
Cleavage of large DNA
molecules by restriction enzymes produces a bewildering array of fragments. How
can the DNA sequence of interest be picked out of a mixture of thousands or
even millions of irrelevant DNA fragments? The answer lies in the use of a
probe, a short piece of ssDNA or RNA, labeled with a radioisotope, such as 32P,
or with a nonradioactive molecule, such as biotin or a fluorescent dye. The
sequence of a probe is complementary to a sequence in the DNA of interest, called
the target DNA. Probes are used to identify which band on a gel or which clone
in a library contains the target DNA, a process called screening.
A. Hybridization of a probe to DNA fragments
The utility of probes
hinges on the phenomenon of hybridization (or annealing) in which a probe
containing a complementary sequence binds a single-stranded sequence of a
target DNA. ssDNA, produced by alkaline denaturation of dsDNA, is first bound
to a solid support, such as a nitrocellulose membrane. The immobilized DNA
strands are prevented from self-annealing but are available for hybridization
to the exogenous, radiolabeled, ssDNA probe. The extent of hybridization is
measured by the retention of radioactivity on the membrane. Excess probe
molecules that do not hybridize are removed by washing the membrane.
B. Synthetic oligonucleotide probes
If the sequence of all
or part of the target DNA is known, short, single-stranded oligonucleotide
probes can be synthesized that are complementary to a small region of the gene
of interest. If the sequence of the gene is unknown, the amino acid sequence of
the protein, the final gene product, may be used to construct a nucleic acid
probe using the genetic code as a guide. Because of the degeneracy of the
genetic code, it is necessary to synthesize several oligonucleotides. [Note:
Oligonucleotides can be used to detect single-base changes in the sequence to
which they are complementary. In contrast, cDNA probes contain many thousands
of bases, and their binding to a target DNA with a single-base change is
unaffected.]
1. Detecting the βS-globin mutation: A synthetic allele-specific
oligonucleotide (ASO) probe can be used to detect the presence of the sickle
cell mutation in the β-globin gene (Figure 33.10). DNA, isolated from
leukocytes and amplified, is denatured and applied to a membrane. A
radiolabeled oligonucleotide probe, complementary to the point mutation (GAG →
GTG, glutamate → valine) at codon 6 in patients with the bS gene, is applied to
the membrane. DNA isolated from a heterozygous individual (sickle cell trait)
or a homozygous patient (sickle cell disease) contains a sequence that is
complementary to the probe, and a double-stranded hybrid forms that can be
detected by electrophoresis. In contrast, DNA obtained from normal individuals
is not complementary at this positon and, therefore, does not form a hybrid
(see Figure 33.10). Use of a pair of such ASO probes (one specific for the
normal allele and one specific for the mutant allele) allows all three possible
genotypes (homozygous normal, heterozygous, and homozygous mutant) to be
distinguished (Figure 33.11). [Note: ASO probes are useful only if the mutation
and its location are known.]
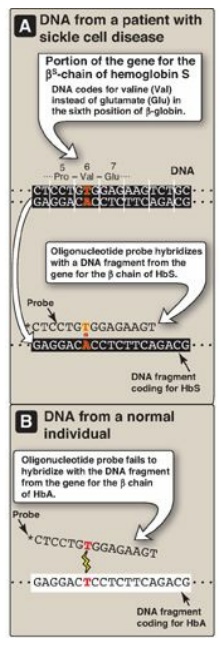
Figure 33.10 Allele-specific
oligonucleotide probe detects hemoglobin (Hb) S allele.
[Note: * indicates 32P radiolabel.] A = adenine; C
= cytosine; G = guanine; T = thymine; Pro = proline.
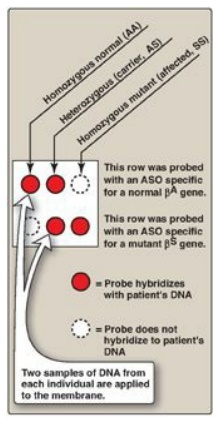
Figure 33.11 Allele-specific
oligonucleotide (ASO) probes used to detect the sickle cell mutation and
differentiate between sickle cell trait and disease.
C. Biotinylated probes
Because the disposal of
radioactive waste is becoming increasingly expensive, nonradiolabeled probes
have been developed. One of the most successful is based on the vitamin biotin,
which can be chemically linked to the nucleotides used to synthesize the probe.
Biotin was chosen because it binds very tenaciously to avidin, a readily
available protein contained in chicken egg whites. Avidin can be attached to a
fluorescent dye detectable optically with great sensitivity. Thus, a DNA
fragment (displayed, for example, by gel electrophoresis) that hybridizes with
the biotinylated probe can be made visible by immersing the gel in a solution
of dye-coupled avidin. After washing away the excess avidin, the DNA fragment
that binds the probe is fluorescent. [Note: Labeled probes can allow detection
and localization of DNA or RNA sequences in cell or tissue preparations, a
process called in situ hybridization (ISH). If the probe is fluorescent, the
technique is called FISH.]
D. Antibodies
If no amino acid
sequence information is available to guide the synthesis of a probe for direct
detection of the DNA of interest, a gene can be identified indirectly by
cloning cDNA in an expression vector that allows the cloned cDNA to translated.
A labeled antibody is used to identify which bacterial protein and, therefore,
contains the cDNA of interest.
Related Topics
