Regulation of Enzyme Activity
| Home | | Biochemistry |Chapter: Biochemistry : Enzymes
The regulation of the reaction velocity of enzymes is essential if an organism is to coordinate its numerous metabolic processes.
REGULATION OF ENZYME ACTIVITY
The regulation of the
reaction velocity of enzymes is essential if an organism is to coordinate its
numerous metabolic processes. The rates of most enzymes are responsive to
changes in substrate concentration, because the intracellular level of many
substrates is in the range of the Km. Thus, an increase in substrate
concentration prompts an increase in reaction rate, which tends to return the
concentration of substrate toward normal. In addition, some enzymes with
specialized regulatory functions respond to allosteric effectors and/or
covalent modification or they show altered rates of enzyme synthesis (or
degradation) when physiologic conditions are changed.
A. Regulation of allosteric enzymes
Allosteric enzymes are
regulated by molecules called effectors that bind noncovalently at a site other
than the active site. These enzymes are almost always composed of multiple
subunits, and the regulatory (allosteric) site that binds the effector is
distinct from the substrate-binding site and may be located on a subunit that
is not itself catalytic. Effectors that inhibit enzyme activity are termed
negative effectors, whereas those that increase enzyme activity are called
positive effectors. Positive and negative effectors can affect the affinity of
the enzyme for its substrate (K0.5), modify the maximal catalytic activity of
the enzyme (Vmax), or both (Figure 5.16). [Note: Allosteric enzymes
frequently catalyze the committed step early in a pathway.]
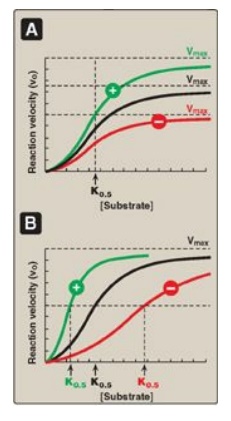
Figure 5.16 Effects of negative 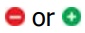 positive effectors on an allosteric enzyme. A. Vmax is
altered. B. The substrate concentration that gives half-maximal velocity (K0.5)
is altered.
positive effectors on an allosteric enzyme. A. Vmax is
altered. B. The substrate concentration that gives half-maximal velocity (K0.5)
is altered.
1. Homotropic effectors: When the substrate itself serves
as an effector, the effect is said to be homotropic. Most often, an allosteric
substrate functions as a positive effector. In such a case, the presence of a
substrate molecule at one site on the enzyme enhances the catalytic properties
of the other substrate-binding sites. That is, their binding sites exhibit
cooperativity. These enzymes show a sigmoidal curve when reaction velocity (vo)
is plotted against substrate concentration ([S]), as shown in Figure 5.16. This
contrasts with the hyperbolic curve characteristic of enzymes following
Michaelis-Menten kinetics, as previously discussed. [Note: The concept of
cooperativity of substrate binding is analogous to the binding of oxygen to
hemoglobin.]
2. Heterotropic effectors: The effector may be different from
the substrate, in which case the effect is said to be heterotropic. For
example, consider the feedback inhibition shown in Figure 5.17. The enzyme that
converts D to E has an allosteric site that binds the endproduct, G. If the
concentration of G increases (for example, because it is not used as rapidly as
it is synthesized), the first irreversible step unique to the pathway is
typically inhibited. Feedback inhibition provides the cell with appropriate
amounts of a product it needs by regulating the flow of substrate molecules
through the pathway that synthesizes that product. Heterotropic effectors are
commonly encountered. For example, the glycolytic enzyme phosphofructokinase-1
is allosterically inhibited by citrate, which is not a substrate for the
enzyme.
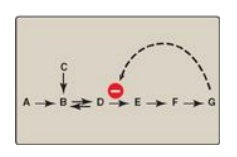
Figure 5.17 Feedback
inhibition of a metabolic pathway.
B. Regulation of enzymes by covalent modification
Many enzymes are
regulated by covalent modification, most often by the addition or removal of
phosphate groups from specific serine, threonine, or tyrosine residues of the
enzyme. Protein phosphorylation is recognized as one of the primary ways in
which cellular processes are regulated. [Note: Protein phosphorylation is
mediated by hormonal signals.]
1. Phosphorylation and dephosphorylation: Phosphorylation reactions are
catalyzed by a family of enzymes called protein kinases that use ATP as the
phosphate donor. Phosphate groups are cleaved from phosphorylated enzymes by
the action of phosphoprotein phosphatases (Figure 5.18).
2. Response of enzyme to phosphorylation: Depending on the specific enzyme,
the phosphorylated form may be more or less active than the unphosphorylated
enzyme. For example, phosphorylation of glycogen phosphorylase (an enzyme that degrades
glycogen) increases activity, whereas phosphorylation of glycogen synthase (an
enzyme that synthesizes glycogen) decreases activity.
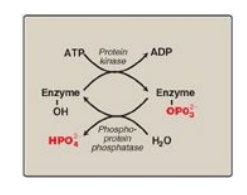
Figure 5.18 Covalent modification by the addition and removal of phosphate groups. [Note: HPO42−may be represented as Pi.]
C. Induction and repression of enzyme synthesis
The regulatory
mechanisms described above modify the activity of existing enzyme molecules.
However, cells can also regulate the amount of enzyme present by altering the
rate of enzyme degradation or, more typically, the rate of enzyme synthesis.
The increase (induction) or decrease (repression) of enzyme synthesis leads to
an alteration in the total population of active sites. Enzymes subject to
regulation of synthesis are often those that are needed at only one stage of
development or under selected physiologic conditions. For example, elevated
levels of insulin as a result of high blood glucose levels cause an increase in
the synthesis of key enzymes involved in glucose metabolism. In contrast,
enzymes that are in constant use are usually not regulated by altering the rate
of enzyme synthesis. Alterations in enzyme levels as a result of induction or
repression of protein synthesis are slow (hours to days), compared with allosterically
or covalently regulated changes in enzyme activity, which occur in seconds to
minutes. Figure 5.19 summarizes the common ways that enzyme activity is
regulated.
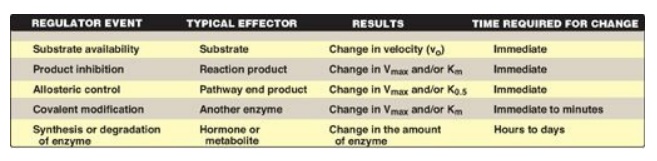
Figure 5.19 Mechanisms for
regulating enzyme activity. [Note: Inhibition by pathway end product is also
referred to as feedback inhibition.]
Related Topics
